Four Cold Responsive Genes That Show Consistent Differential Expression between the Japonica and Indica subspecies of rice
*Corresponding Author(s):
Donghai MaoInstitute Of Subtropical Agriculture, Chinese Academy Of Sciences, Beijing, China
Tel:+ 86 73184619748,
Email:donghai.mao@isa.ac.cn
Abstract
Rice (Oryza sativa L.) is highly sensitive to cold stress during the seedling stage and of the two cultivated subspecies, japonica rice varieties tend to be more tolerant to low temperatures than are indica rice varieties. In this study, we performed comparative transcriptome analysis and found 171 genes that respond to cold under four different conditions in the two rice subspecies. qRT-PCR assays and natural variation analysis were further carried out to identify the differentially-expressed Cold Responsive genes (CORs) involved in the difference in cold tolerance between the japonica and indica rice subspecies. We identified four genes, OsDREB1A, OsABCB5, OsPHIL2 and OsREM4. 1 that showed consistently different expression patterns in the two rice subspecies in response to low temperature. Based on the results of nature variation analysis of these four genes, we predicted they respond to cold through the CBF/DREB1or plant hormone pathway, respectively. The four differentially expressed genes may be key genes lead to different cold tolerance between two rice subspecies. These results not only contribute to a comprehensive understanding of the differences in transcriptome levels of low temperature response genes between rice subspecies, but also provide candidate genes for molecular breeding of cold-tolerant rice cultivars.
Keywords
INTRODUCTION
Rice (Oryza sativa L.) is the most important staple crop that feeds nearly half of the world’s population. Because originated in tropical or subtropical areas, rice is sensitive to low temperatures throughout its entire life cycle. Cold stress can inhibit photosynthesis by reducing the chlorophyll concentration, inducing the accumulation of Reactive Oxygen Species (ROS) and by causing severe damage to various cellular components including membrane lipids, structural proteins and enzymes [1-3]. Low temperatures have the potential to negatively affect the growth and development of rice plants during any developmental stage, from germination to grain filling, resulting in large losses in rice production [4].
Plants have developed several strategies to respond to low temperatures that are known collectively as ‘cold acclimation’ [5]. Numerous studies of cold acclimation have been reported in recent years, and they showed that many pathways participate in the cold-response process [6,7]. The Dehydration-Responsive Element Binding protein (DREB1)/C-repeat binding factor (CBF)-dependent cold-responsive pathway is the most well-known among all the pathways [8]. Another pathway with a central role in the rice response to cold stress is the MYBS3-dependent pathway. Interestingly, DREB1A responds early and transiently, while MYBS3 responds relatively slowly to cold stress in rice, suggesting that theMYBS3-dependent pathway is important for long-term adaptation to persistent cold stress [9]. In addition to these two pathways, the plant hormone systems play an indispensable role in cold stress tolerance [10]. For example, Jasmonate (JA) positively regulates cold tolerance in plants because it can positively modulate the CBF-dependent or –independent pathway, leading to accumulation of cryoprotective compounds such as polyamine, glutathione and anthocyanins [11]. DELLA proteins, the key components of the Gibberellic Acid (GA)-signaling pathway, are involved in the CBF1-mediated cold stress response [12]. The cross-talk between Cytokinin (CK) and Abscisic Acid (ABA) are also important in cold stress signaling. Components of CK signaling, Histidine Kinases (AHKs) and the cold-inducible A-type ARRs, play negative regulatory roles in cold stress response via inhibition of the ABA response [13]. Components of Brassinosteroid (BR) and ethylene signaling were also found to modulate freezing tolerance in Arabidopsis plants [14,15].
There are two major subspecies of rice, Oryza sativa ssp. Indica and Oryza sativa ssp. japonica. Generally, japonica rice varieties are more tolerant to low temperatures than varieties of indica rice [16]. Therefore, cold tolerance in indica rice can be improved by introducing genes from japonica rice through molecular breeding. Recent advances in next generation sequencing technology provide a powerful strategy for discovering new genes associated with cold tolerance in rice [17-21]. In this study, we combined transcriptome sequencing with genome re-sequencing to identify the Cold-Responsive-genes (CORs) that are differentially-expressed between the two rice subspecies. Using this approach, we found four COR genes that are differentially-expressed between indica and japonica varieties. Moreover, these genes show important natural variation between the two subspecies which predicted to result in changes of cold responsive cis-elements. Therefore, these CORs may be useful for improving cold tolerance in rice through molecular breeding.
RESULTS
Global mRNA expression profiles in rice seedlings in response to cold stress
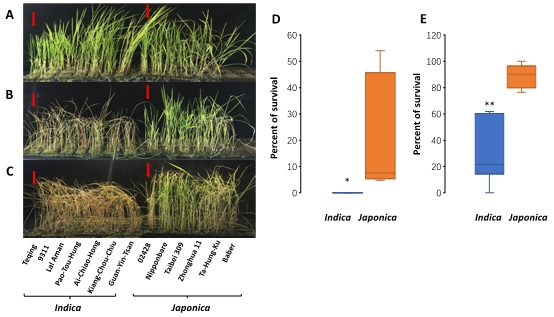
Figure S1: Phenotypes of the seedlings of the indica and japonica rice cultivars before (A) and after exposure at 4°C for 4 days (B) and 10oC for 5 days(C).Teqing and 02428 are indicated with red arrows). Graphical representation of the relative survival rates of 7 indica and 6 japonica seedlings following exposure to cold stress at 4°C for 4 days (D) or10°C for 5 days (E). Mean values with one or two asterisks were found to be significantly different by Student's t test (*P≤0.05; **P≤0.01; n≥6).
|
Samples |
Raw Tags |
Distinct Tags |
Clean Tags |
Distinct Clean Tags |
Map to Gene Tags |
Unique Map to Gene Tags |
Map to Genome Tags |
Unknown Tags |
|
J0 |
5858191 |
251288 |
5721402 |
115619 |
4967023(86.81%) |
2821188 (49.31%) |
473352 (8.27%) |
281027 (4.91%) |
|
4J3 |
5846716 |
284227 |
5688748 |
126879 |
4896262(86.07%) |
2747337 (48.29%) |
530238 (9.32%) |
262248 (4.61%) |
|
4J24 |
5913697 |
294409 |
5745004 |
127466 |
4829848(84.07%) |
2780519 (48.40%) |
704593 (12.26%) |
210563 (3.67%) |
|
10J3 |
6215714 |
301590 |
6048834 |
136023 |
4993999(82.56%) |
2788766 (46.10) |
576051 (9.52%) |
478784 (7.92%) |
|
10J24 |
5867356 |
280913 |
5714812 |
128893 |
4730647(82.78%) |
2552274 (44.70%) |
496393 (8.68%) |
487862 (8.54%) |
|
I0 |
5972875 |
241437 |
5841753 |
111174 |
4865519(83.29%) |
2747870 (47.04%) |
548967 (9.40%) |
427267 (7.31%) |
|
4I3 |
5800974 |
259563 |
5657990 |
117336 |
4638484(81.98%) |
2574611 (45.50%) |
566161 (10.01%) |
453345 (8.01%) |
|
4I24 |
6145845 |
272766 |
5996689 |
124950 |
4889203(81.53%) |
2810809 (46.87%) |
755255 (12.59%) |
352231 (5.87%) |
|
10I3 |
6104075 |
289808 |
5941508 |
128383 |
4916259(82.74%) |
2781934 (46.82%) |
607244 (10.22%) |
418005 (7.04%) |
|
10I24 |
5882748 |
292303 |
5713363 |
124585 |
4710977(82.46%) |
2603310 (45.57%) |
5226813 (9.22%) |
475573 (8.32%) |
Table 1: Numbers of sequence tags in the 10 transcriptome libraries from rice cultivars Teqing and 02428 exposed to low temperatures.
Identification of COR genes in the indica and japonica rice cultivars
Gene ontology and pathway enrichment analysis of the rice CORs

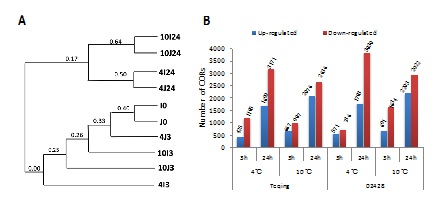
Figure 2: Gene Ontology (GO) analysis of COR genes in the “molecular function” (A), “biological process” (B) and “cell component” (C) GO classes in Teqing and 02428 at 4°C and 10°C for 3hr and 24hr. The y-axis and x-axis indicate the percentage of genes in a GO category and the names of the clusters, respectively. The numbers before the letter (I or J) indicate the temperatures, the numbers after the letter (I or J) indicate the hours of exposure, J and I mean Teqing and 02428, respectively. For example, ‘CORs in 4I3’ means differentially expressed genes in Teqing between before (0hr) and after (3hr) cold treatment (4I3 vs I0, |log2Ratio| ?1 and FDR ≤ 0.001).
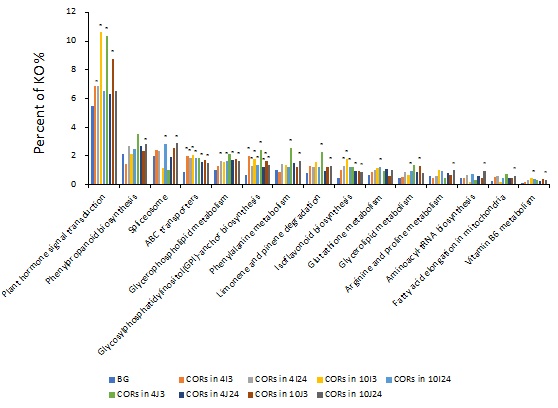
BG refers to all the genes in the background. ‘CORs in 4I3’ means differentially expressed genes in Teqing between before (0hr) and after (3hr) cold treatment (4I3 vs I0, |log2Ratio| ≥1 and FDR ≤ 0.001). By analogy, the numbers before the letter (I or J) indicate the temperatures, the numbers after the letter (I or J) indicate the hours of exposure; J and I mean Teqing and 02428, respectively.* indicates significantly enriched KEGG pathways with correct P value ≤0.05.
Identification of consistent differentially-expressed COR genes in multiple rice cultivars
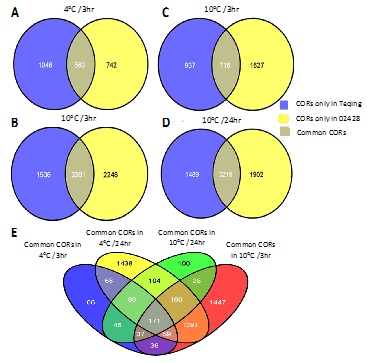
Figure 4: Venn diagrams showing the shared COR genes in Teqing and 02428 after exposure to cold treatments. (A) 563 COR genes respond to cold in both cultivars at 4°C for 3 hours; (B) 3361 COR genes respond to cold in both cultivars at 4°C for 24 hours; (C) 716 COR genes respond to cold in both cultivars at 10°C for 3 hours; (D) 3219 COR genes respond to cold in both cultivars at 10°C for 24 hours (CORs only in Teqing are shown in blue circle with the number written in white, CORs only in 02428 are shown in yellow circle with the number written in black, CORs shared by both cultivars are shown in center part of the Venn diagrams with the number written in white). (E) among all the common CORs we get from picture A, B, C and D, 171 COR genes respond to cold in both cultivars in all these cold treatments (Words above the pictures indicate the source of CORs. For example, common CORs in 4°C/3hr means CORs shared by Teqing and 02428, when they are treated in 4°C for 3 hours).
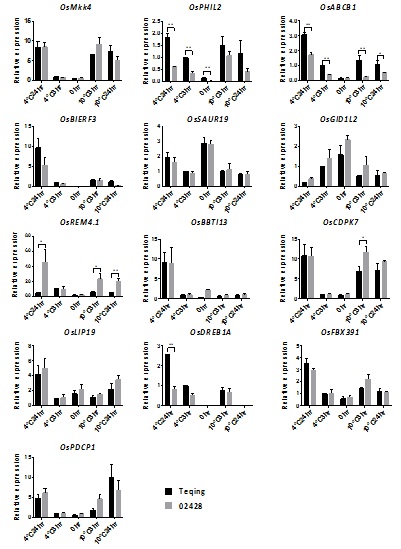
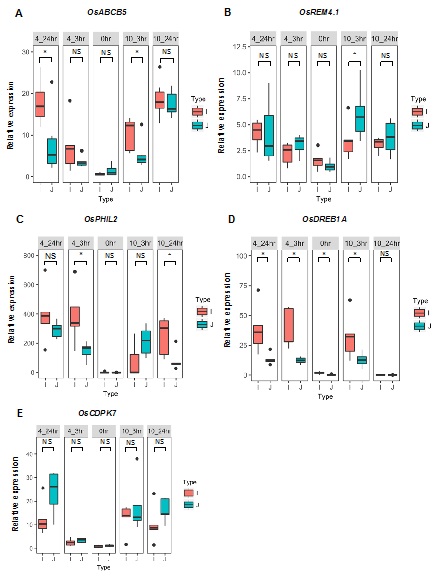
Figure 5: qRT-PCR analysis of 13 selected COR genes identified by RNA-seq in leaves of Teqing and 02428 plants before (0hr) and after 3 h or 24 h of cold treatment at 4°C or 10°C. The data of expression levels were separately compared with the Teqing sample before cold treatment (0hr). OsActin was used as an internal standard (Student’s t test, *P≤0.05, **P≤0.01, n=3).
To determine whether the observed differences in expression for the five genes are associated with cold tolerance in other cultivars of the two subspecies, we included an additional five cold tolerant japonica rice cultivars (Nipponbare, Ta Hung Ku, Taibei309, Baber, Zhonghua 11) and five cold sensitive indica rice cultivars (9311, Lal-Aman, Pao-Tou-Hung, Ai-Chiao-Hong, Guan-Yin-Tsan) to the study. Details of the cold responses of these 10cultivars are shown in figure S1. All of the japonica rice cultivars showed significantly stronger cold tolerance than the indica cultivars, as expected. We then we assayed the expression levels of the five genes (OsABCB5, OsPHIL2, OsDREB1A, OsCDPK7 and OsREM4.1) in the 10 cultivars. Among these five genes, the expression profiles of OsABCB5, OsPHIL2, OsDREB1A and OsREM4. 1still showed clear distinctions between the indica and japonica subspecies, while OsCDPK7 did not show the indica-japonica difference in the multi-cultivar comparison (Figure 6). In the initial comparison between Teqing and 02428, OsABCB5, OsPHIL2 and OsDREB1Awere expressed at higher levels in Teqing than in 02428; similarly, in the comparison of the five cultivar pairs, the expression of the three genes was also higher in indica than in japonica. OsREM4.1 showed higher expression levels in the japonica rice cultivars, confirming the results from the comparison of Teqing and 02428. After cold treatment, expression of OsABCB5, OsPHIL2, OsDREB1A and OsREM4. 1 was significantly higher than in the control, and the expression of OsPHIL2 was up-regulated >400-fold after 4°C treatment in the indica cultivars.
Analysis of natural variation in these COR genes in multiple rice cultivars
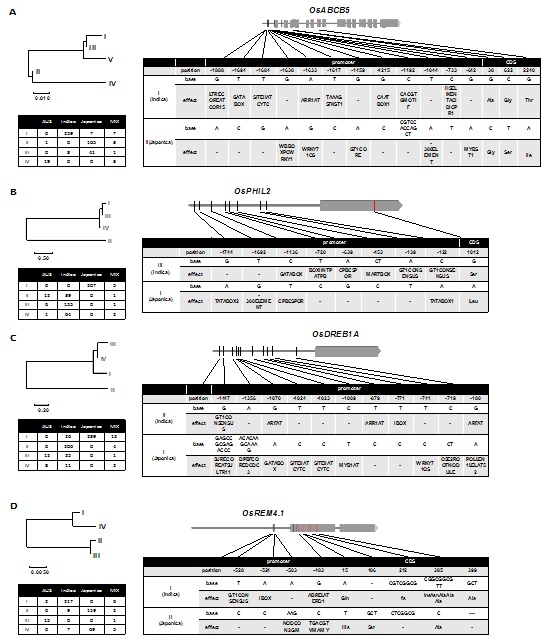
Figure 7: Schematic diagrams showing the position of SNP/Indels in the2.0-kb promoter regions and Coding Regions (CDS) of the OsABCB5 (A), OsPHIL2 (B), OsDREB1A (C) and OsREM4.1(D) genes. The phylogenetic tree was generated by MEGA5.0 software using neighbor joining method, with the segment length of interior branches indicating bootstrap values (1000 replications). The scale shows nucleotide substitutions per site. In gene schematic diagrams, gray bars indicate exons, and black lines indicate introns, the red vertical lines indicate the SNPs in coding regions while the black vertical lines indicate the SNPs in promoter regions. In the left table, Aus, Indica, Japonica and MIX represent the ecotypes of cultivated rice, such as aus, indica, japonica and mixture of other ecotypes, and the numbers in each block show the number of varieties. In the right table, the position of TSS is 0, negative and positive numbers indicate the positions of polymorphic sites in the promoter and coding region, respectively.
Based on nucleotide polymorphisms, we could divide the sequences of the 575 lines into four haplotypes for OsABCB5; among these, HapII was the most represented in japonica lines, and Hap I was the most represented in indica lines. There were 12 SNP/Indel variations identified in the OsABCB5 promoter region between indica and japonica lines. Of these, four were predicted to be located within low temperature responsive cis-elements. For example, Hap II and Hap I differed in the nucleotides located at positions -642 G/A, -1633 A/G, -1638 G/A, and -1808 G/A upstream of the start codon; these four SNPs were related to theMYBST1, WRKY71OS/ARR1AT, WBBOXPCWRKY1 and LTRECOREATCOR15motifs, respectively, that are predicted to function in the cold response.
DISCUSSION
The japonica subspecies of Oryza sativa has adapted to a temperate climate, while the indica subspecies has adapted to tropical and subtropical environments. As a result, japonica rice cultivars are generally more tolerant to cold stress than indica rice cultivars (Figure S1). In our study, we used comparative transcriptome analysis on two rice cultivars to identify a group of COR genes in these two cultivars. From the CORs, we further identified four key genes, OsABCB5, OsPHIL2, OsDREB1A and OsREM4.1, that showed ubiquitous and distinctive activity in indica and japonica cultivars in an integrative study using KEGG and GO annotations, Venn diagram analysis, qRT-PCR, and natural variation analysis.
The DREB1/CBF pathway is a well-known cold-responsive pathway in plants [23]. The pathway includes the CBF1, CBF2 and CBF3 genes, which are also known as DREB1B, DREB1C and DREB1A, respectively, in Arabidopsis [24]. However, not all three transcription factors act as positive regulators in the CBF/DREB1 pathway. In Arabidopsis, freezing tolerance of the cbf2 mutant is enhanced by up-regulating the expression of CBF1/DREB1B and CBF3/DREB1A, which means that CBF2/DREB1C plays a negative role in the cold response [25]. OsDREB1A has been reported previously to be induced by cold stress [26,27]. Consistent with these results, we found that the expression of OsDREB1Aincreased after cold treatment. Lourenco et al., have claimed that in OsHOS1 knockdown plants, the reduced expression of OsHOS1 promoted the accumulation of OsICE1 protein and resulted in the up-regulation of OsDREB1A. However, the transgenic plants did not show increased cold tolerance [28]. In our study, the expression level of OsDREB1A was higher in indica rice compared to japonica rice. Therefore, we speculate that OsDREB1Amay play a negative role in cold signaling. In our natural variation analysis, we discovered a SNP related to MYB in the promoter region of OsDREB1Ain HapI (japonica). Interestingly, a MYB-like transcription factor has been reported to be involved in the cold regulation of CBF genes, and MYB15 over expression reduced the expression of CBF genes under cold treatment [29]. Thus, the MYB negative regulation of OsDREB1A may result in the stronger cold tolerance observed in japonica rice cultivars.
Among the four genes we screened, OsREM4.1 is related to the plant hormone ABA, which is known as a cold stress-responsive hormone. Two SNP variations within ABA responsive or MYB elements were located within the promoter region of OsREM4.1. Interestingly, OsREM4.1 was previously characterized and found to be up-regulated by ABA signaling; the OsREM4.1 protein can bind to OsSERK1 and inhibit the formation and activation of the OsBRI1-OsSERK1 receptor complex, which is crucial for BR signaling [30]. In Arabidopsis thaliana, application of exogenous ABA can enhance cold tolerance and induce the expression of several COR genes, including RAB18, RD29A and KIN2 [31,32]. Several studies showed that plants treated with BR grew better at low temperature compared to optimal conditions [33]. In addition, Li et al., described two key components of BR signaling, BIN2 and BZR1, regulate plant freezing tolerance; BIN2 and its homologs play negative roles, while BZR1 plays a positive role in the cold response [14]. In our study, OsREM4.1hada higher level of expression in japonica rice, so we predicted that OsREM4.1 may play a positive role in regulating cold tolerance via crosstalk between the ABA and BR pathways.
Another gene we found that is associated with the BR pathway is OsPHIL2, which encodes a Phosphate-Induced protein (PHI). Interestingly, two BR-response genes in Arabidopsis, EXORDIUM (EXO) and EXORDIUM-LIKE1 (EXL1), were previously reported to have the same PHI conserved region [34]. Expression of the EXO gene is up-regulated by BR and EXL1 has a similar expression pattern to EXO in Arabidopsis in that it can be induced by BR [35,36]. We therefore predicted that OsPHIL2 may be similarly connected to BR signaling pathway. Considering that OsPHIL2 is expressed at higher levels in indica cultivars when exposed to low temperature, OsPHIL2 may negatively regulate cold stress via BR signaling.
ABC transporters constitute one of the largest protein families, and are present in organisms ranging from bacteria to humans [37]. In plants, ABC proteins were originally identified as transporters involved in the final detoxification process [38]. OsABCB5 is predicted to encode an MDR-like ABC transporter. There are several studies that have reported that MDR-like ABC transporters are involved in the auxin signaling pathway. Noh et al., showed that two MDR-like genes, At MDR1 and AtPGP1, are required for auxin transport and auxin-mediated development in Arabidopsis. Also, mutation of two related MDR-like genes (MDR1 and PGP1) resulted in reduced polar auxin transport [39]. Another study found that the Arabidopsis MDR-like ABC transporter4 (AtPGP4) is involved in auxin-mediated root development [40]. Although the role of auxin in cold stress remains unclear, there are reports that potentially link cold stress to auxin by showing that cold stress inhibits the inflorescence gravity response in Arabidopsis thaliana [41,42]. An early study demonstrated that temperature affects the speed of exogenous auxin transport in a variety of plant species [43]. Considering that auxin is at the center of hormonal crosstalk, auxin may mediate the response to cold by interacting with other hormones [44]. These results indicate that OsABCB5may play a crucial role in the cold response in rice seedlings by transporting auxin.
Based on the expression profiles and predicted pathways of the four key genes involved in the cold response in rice, we can conduct further functional studies to determine the biological and biochemical functions of the proteins encoded by these genes.
MATERIALS AND METHOD
Plant materials and Evaluation of Phenotypes
RNA isolation
Library construction and Illumina cDNA sequencing
Identification of COR genes
Quantitative reverse transcription polymerase chain reaction (qRT-PCR) analysis of gene expression
|
Locus |
Gene |
Annotation |
Forward |
Reverse |
|
LOC_Os02g52010 |
OsPHIL2 |
Phosphate-induced protein 1 conserved region domain containing protein |
CACCATCAACCAGCTGTACC |
ATCCCGCCCTTCTTGGGCTT |
|
LOC_Os01g50100 |
OsABCB1 |
ABC transporter, ATP-binding protein |
GAGAGGAAACCAGAGATAG |
TTCGCCAGACTGTGGATCGTA |
|
LOC_Os07g38170 |
OsREM4.1 |
remorin |
AGATTGTCATCAGCACCG |
GTAGGGAAGAGCTCACTT |
|
LOC_Os09g35030 |
OsDREB1A |
dehydration-responsive element-binding protein |
TTCGAACTGGACGTCCTGAGT |
TAGTAGCTCCAGAGTGGGA |
|
LOC_Os02g43790 |
OsBIERF3 |
ethylene-responsive transcription factor |
TTAATCCGGCGTCGAGAGA |
AAGCTTGAGCTCCGGCAGTA |
|
OsSAUR19 |
OsSAUR19 - Auxin-responsive SAUR gene family member |
TTGCAAGGAGGGGAGCGAAGAA |
TTGAAGTACACCACCGGCA |
|
|
LOC_Os09g28740 |
Gibberellin receptor GID1L2 |
AAGTACCACGACTACCTGA |
TTCTTGGCCACCCAGTTCA |
|
|
LOC_Os03g60840 |
BBTI13 - Bowman-Birk type bran trypsin inhibitor precursor |
CTCTGTTCTTGGCCTTTGT |
CGCAGTTGTCGCAGCA |
|
|
OsCDPK7 |
CAMK includes calcium/calmodulin depedent protein kinases |
ACACCGAGATTCGTGATC |
TGCAAGCTTGCTGCAGTT |
|
|
LOC_Os03g55240 |
cytochrome P450 |
ATGTTGCTCATCAACGCGTA |
CAGTCGAAGCACTGGATCA |
|
|
LOC_Os10g37830 |
OsFBX391 - F-box domain containing protein |
GAGTTTGCAGCACTGAGCAT |
ATCTGCGCCATCGAGAACTTCA |
|
|
LOC_Os08g37670 |
Plastocyanin-like domain containing protein |
CTTGGCCAGAACTACGATAC |
TGGTCGAGCTGATCGAGTT |
|
|
LOC_Os02g54600 |
OsMKK4 |
TE_MEK_ste7_MAP2K.5 - STE kinases include homologs to sterile 7, sterile 11 and sterile 20 from yeast |
ACATCAAGCCATCCAACCT |
TAGAACTCGAGAATGCTCA |
|
LOC_Os03g50885 |
OsActin |
ACACCGGTGTCATGGTCGG |
ACACGGAGCTCGTTGTAGAA |
Table S1: Sequences of the oligonucleotide primers used in the qRT-PCR assays.
Haplotype analysis and SNP discovery of the four key COR genes
AUTHOR CONTRIBUTION
DM conceived and designed the experiments. XH and DM performed the experiments. XH and YT analyzed the data, and XH wrote the manuscript.
ACKNOWLEDGEMENT
We thank Dr. Caiyan Chen for his help in conceiving the project. This work was supported by grants from the National Natural Science Foundation of China (31101211, 31371603).
REFERENCES
- Aghaee A, Moradi F, Zare-Maivan H, Zarinkamar F, Irandoost HP, et al. (2011) Physiological Responses of Two Rice (Oryza Sativa L.) Genotypes to Chilling Stress at Seedling Stage. African J Biotechnol 10: 7617-7621.
- Song SY, Chen Y, Chen J, Dai XY, Zhang WH (2011) Physiological Mechanisms Underlying OsNAC5-Dependent Tolerance of Rice Plants to Abiotic Stress. Planta 234: 331-345.
- Uemura M, Joseph RA, Steponkus PL (1995) Cold Acclimation of Arabidopsis Thaliana (Effect on Plasma Membrane Lipid Composition and Freeze-Induced Lesions). Plant Physiol 109: 15-30.
- Ye H, Du H, Tang N, Li X, Xiong L (2009) Identification and Expression Profiling Analysis of TIFY Family Genes Involved in Stress and Phytohormone Responses in Rice. Plant Mol Biol 71: 291-305.
- Thomashow MF (1999) Plant Cold Acclimation: Freezing Tolerance Genes and Regulatory Mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50: 571-599.
- Rihan HZ, Al-issawi M, Fuller MP (2017) Advances in Physiological and Molecular Aspects of Plant Cold Tolerance. J PLANT Interact 12: 143-157.
- Shi Y, Ding Y, Yang S (2015) Cold Signal Transduction and Its Interplay with Phytohormones during Cold Acclimation. Plant Cell Physiol 56: 7-15.
- Ito Y, Kurata N (2006) Identification and Characterization of Cytokinin-Signalling Gene Families in Rice. Gene 382: 57-65.
- Su CF, Wang YC, Hsieh TH, Lu CA, Tseng TH, et al. (2010) A Novel MYBS3-Dependent Pathway Confers Cold Tolerance in Rice. Plant Physiol 153: 145-158.
- Kurepin LV, Dahal KP, Savitch LV, Singh J, Bode R, et al. (2013) Role of CBFs as Integrators of Chloroplast Redox, Phytochrome and Plant Hormone Signaling during Cold Acclimation. Int J Mol Sci 14: 12729-12763.
- Hu Y, Jiang L, Wang F, Yu D (2013) Jasmonate Regulates The Inducer Of Cbf Expression-C-Repeat Binding Factor/Dre Binding Factor1 Cascade and Freezing Tolerance in Arabidopsis. Plant Cell 25: 2907-2924.
- Achard P, Gong F, Cheminant S, Alioua M, Hedden P, et al. (2008) The cold-inducible CBF1 factor-dependent signaling pathway modulates the accumulation of the growth-repressing DELLA proteins via its effect on gibberellin metabolism. Plant Cell 20: 2117-2129.
- Jeon J, Kim NY, Kim S, Kang NY, Novák O, et al. (2010) A Subset of Cytokinin Two-Component Signaling System Plays a Role in Cold Temperature Stress Response in Arabidopsis. J Biol Chem 285: 23371-23386.
- Li H, Ye K, Shi Y, Cheng J, Zhang X,et al. (2017) BZR1 Positively Regulates Freezing Tolerance via CBF-Dependent and CBF-Independent Pathways in Arabidopsis. Mol Plant 10: 545-559.
- Zhang Z, Huang R (2010) Enhanced Tolerance to Freezing in Tobacco and Tomato Overexpressing Transcription Factor TERF2/LeERF2 Is Modulated by Ethylene Biosynthesis. Plant Mol Biol 73: 241-249.
- Lv Y, Guo Z, Li X, Ye H, Li X, et al. (2016) New Insights into the Genetic Basis of Natural Chilling and Cold Shock Tolerance in Rice by Genome-Wide Association Analysis. Plant Cell Environ 39: 556-570.
- Bai B, Wu J, Sheng WT, Zhou B, Zhou LJ, et al. (2015) Comparative Analysis of Anther Transcriptome Profiles of Two Different Rice Male Sterile Lines Genotypes under Cold Stress. Int J Mol Sci 16: 11398-11416.
- Shen C, Li D, He R, Fang Z, Xia Y, et al. (2014) Comparative Transcriptome Analysis of RNA-Seq Data for Cold-Tolerant and Cold-Sensitive Rice Genotypes under Cold Stress. J Plant Biol 57: 337-348.
- Dametto A, Sperotto RA, Adamski JM, Blasi ÉA, Cargnelutti D, et al. (2015) Cold Tolerance in Rice Germinating Seeds Revealed by Deep RNAseq Analysis of Contrasting Indica Genotypes. Plant Sci 238: 1-12.
- Byun MY, Cui LH, Lee J, Park H, Lee A, et al. (2018) Identification of Rice Genes Associated With Enhanced Cold Tolerance by Comparative Transcriptome Analysis With Two Transgenic Rice Plants Overexpressing DaCBF4 or DaCBF7, Isolated From Antarctic Flowering Plant Deschampsia antarctica. Front Plant Sci 9: 601.
- Sperotto RA, de Araújo Junior AT, Adamski JM, Cargnelutti D, Ricachenevsky FK, et al. (2018) Deep RNAseq Indicates Protective Mechanisms of Cold-Tolerant Indica Rice Plants during Early Vegetative Stage. Plant Cell Rep 37: 347-375.
- Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, et al. (2013) Improvement of the Oryza Sativa Nipponbare Reference Genome Using next Generation Sequence and Optical Map Data. Rice 6: 4.
- Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, et al. (2006) Functional Analysis of Rice DREB1/CBF-Type Transcription Factors Involved in Cold-Responsive Gene Expression in Transgenic Rice. Plant Cell Physiol 47: 141-153.
- Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, et al. (1998) Two Transcription Factors, DREB1 and DREB2, with an EREBP/AP2 DNA Binding Domain Separate Two Cellular Signal Transduction Pathways in Drought- and Low-Temperature-Responsive Gene Expression, Respectively, in Arabidopsis. Plant Cell 10: 1391-1406.
- Novillo F, Alonso JM, Ecker JR, Salinas J (2004) CBF2/DREB1C is a negative regulator of CBF1/DREB1B and CBF3/DREB1A expression and plays a central role in stress tolerance in Arabidopsis. Proc Natl Acad Sci USA 101: 3985-3990.
- Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, et al. (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33: 751-763.
- Challam C, Ghosh T, Rai M, Tyagi W (2015) Allele Mining across DREB1A and DREB1B in Diverse Rice Genotypes Suggest a Highly Conserved Pathway Inducible by Low Temperature. J Genet 94: 231-238.
- Lourenço T, Sapeta H, Figueiredo DD, Rodrigues M, Cordeiro A, et al. (2013) Isolation and characterization of rice (Oryza sativa L.) E3-ubiquitin ligase OsHOS1 gene in the modulation of cold stress response. Plant Mol Biol 83: 4-5.
- Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, et al. (2006) A R2R3 Type MYB Transcription Factor is Involved in the Cold Regulation of CBF Genes and in Acquired Freezing Tolerance. J Biol Chem 281: 37636-37645.
- Gui J, Zheng S, Liu C, Shen J, Li J, et al. (2016) OsREM4.1 Interacts with OsSERK1 to Coordinate the Interlinking between Abscisic Acid and Brassinosteroid Signaling in Rice. Dev Cell 38: 201-213.
- Kurkela S, Borg-Franck M (1992) Structure and expression of kin2, one of two cold- and ABA-induced genes of Arabidopsis thaliana. Plant Mol Biol 19: 689-692.
- Lång V, Palva ET (1992) The Expression of a Rab-Related Gene, Rab18, Is Induced by Abscisic Acid during the Cold Acclimation Process of Arabidopsis Thaliana (L.) Heynh. Plant Mol Biol 20: 951-962.
- Kamuro Y, Takatsuto S (1991) Capability for and Problems of Practical Uses of Brassinosteroids. Acs Symp Ser 292-297.
- Schröder F, Lisso J, Müssig C (2012) Expression Pattern and Putative Function of EXL1 and Homologous Genes in Arabidopsis. Plant Signal Behav 7: 22-27.
- Coll-Garcia D, Mazuch J, Altmann T, Müssig C (2004) EXORDIUM Regulates Brassinosteroid-Responsive Genes. FEBS Lett 563: 82-86.
- Schröder F, Lisso J, Lange P, Müssig C (2009) The Extracellular EXO Protein Mediates Cell Expansion in Arabidopsis Leaves. BMC Plant Biol 12: 1-12.
- Henikoff S, Greene EA, Pietrokovski S, Bork P, Attwood TK, et al. (1997) Gene Families: The Taxonomy of Protein Paralogs and Chimeras. Science 278: 609-614.
- Martinoia E, Grill E, Tommasini R, Kreuz K, Amrhein N, (1993) ATP-Dependent Glutathione S-Conjugate “export” Pump in the Vacuolar Membrane of Plants. Nature 364: 247-249.
- Noh B, Bandyopadhyay A, Peer WA, Spalding EP, Murphy AS (2003) Enhanced Gravi- and Phototropism in Plant Mdr Mutants Mislocalizing the Auxin Efflux Protein PIN1. Nature 423: 999-1002.
- Santelia D, Vincenzetti V, Azzarello E, Bovet L, Fukao Y, et al. (2005) MDR-like ABC Transporter AtPGP4 Is Involved in Auxin-Mediated Lateral Root and Root Hair Development. FEBS Lett 579: 5399-5406.
- Fukaki H, Fujisawa H, Tasaka M (1996) Gravitropic Response of Inflorescence Stems In Arabidopsis thaliana. Plant Physiol 110: 933-943.
- Wyatt SE, Rashotte AM, Shipp MJ, Robertson D, Muday GK (2002) Mutations in the Gravity Persistence Signal Loci in Arabidopsis Disrupt the Perception and/or Signal Transduction of Gravitropic Stimuli. Plant Physiol 130: 1426-1435.
- Morris DA (1979) The Effect of Temperature on the Velocity of Exogenous Auxin Transport in Intact Chilling-Sensitive and Chilling-Resistant Plants. Planta 146: 603-605.
- Rahman A (2013) Auxin: A Regulator of Cold Stress Response. Physiol Plant 147: 28-35.
- Yamaji N, Xia J, Mitani-Ueno N, Yokosho K, Feng Ma J (2013) Preferential delivery of zinc to developing tissues in rice is mediated by P-Type heavy metal ATPase OsHMA2. Plant Physiol 162: 927-939.
- Mao D, Yu L, Chen D, Li L, Zhu Y, et al. (2015) Multiple Cold Resistance Loci Confer the High Cold Tolerance Adaptation of Dongxiang Wild Rice (Oryza Rufipogon) to Its High-Latitude Habitat. Theor Appl Genet 128: 1359-1371.
- Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) AgriGO: A GO Analysis Toolkit for the Agricultural Community. Nucleic Acids Res 38: 64-70.
- Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, et al. (1999) KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Research 28: 27-30.
- Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, et al. (2008) KEGG for Linking Genomes to Life and the Environment. Nucleic Acids Res 36: 480-484.
- Tan L, Zhu Y, Fan T, Peng C, Wang J, et al. (2019) OsZIP7 functions in xylem loading in roots and inter-vascular transfer in nodes to deliver Zn/Cd to grain in rice. Biochem Biophys Res Commun 512: 112-118.
Citation: Mao D (2019) Four Cold Responsive Genes That Show Consistent Differential Expression between the Japonica and Indica subspecies of rice. J Genet Genomic Sci 4: 011.
Copyright: © 2019 Donghai Mao, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.