
S-IgG Assay is a Good Replacement of the High Risk, Time Costly Virus Neutralization Assay
*Corresponding Author(s):
Xiang YangLeide Biosciences Co., Ltd, China
Email:yangx@leidebio.com
Jie Wu
The Guangdong Provincial Center For Disease Control And Prevention, Guangzhou, Guangdong, China
Email:771276998@qq.com
Abstract
Coronavirus disease (COVID-19) is caused by a newly discovered virus, SARS-COV-2. About 21 million people were infected and 766 thousand confirmed death. The only hope left is vaccine. Whether a vaccine could work or not is a problem waiting for answer. To find the best method that could predict the protective immunity after SARS-COV-2 virus infection, a comparative study of one virus neutralizing experiment and four ELISA kits was conducted. In one hospital from164 Covid-19 patients that were discharged, 97.56% serum had neutralization activity and 93.29% had anti-S IgG. In another group of 184 persons that had closely contacted with covid-19 patients, 35.9% had neutralization activity and 34.7% had anti-S IgG. TheS- IgG assay shows 93.5% agreement with the virus neutralization experiment, which indicates that more than 90% SARS-COV-2 virus neutralization activity is related to the S protein IgG. S-IgG assay could be used to replace the high risk, time costly virus neutralization assay.
Keywords
COVID-19; Immunity; Pandemic
INTRODUCTION
While the worldwide spreading of SARS-COV-2 virus continues, the appearance of population immunity is expected after infection whether symptomatic or not. There are two options for preventing COVID-19 outbreaks. One is the social segregation model adopted by China. Everyone was required to stay at home for one to two months. Another one is the herd immunity model adopted by some European countries [1]. Not long ago, British Prime Minister Boris Johnson (healed after being infected with this virus) and his government scientific adviser publicly announced that herd immunity may be the ultimate methods to end this pandemic. But if the battle will be won by spreading immunity, which method could be used to detect the immunity after infection? Can we issue a safe passport while this immunity tests are positive? These are questions without an answer yet. The search for a rapidly available, cost-sensitive and accurate test remains active.
The sensitivity of virus nucleic acid detection is affected by sampling, operation, and the quality of detection reagents. Only about 40-70% of infected people test positive [2]. Especially those with mild and asymptomatic infections, which are difficult to be monitored [3]. Moreover, the detection of viral nucleic acid can only prove the presence of the virus, and does not necessarily reflect the patient's immune status. Immunity to viruses is divided into cellular immunity and humoral immunity. Cellular immunity is composed of memory T cells, NK cells and macrophages and has not been studied in this article. Our research focuses on serological antibody testing and virus neutralization.
The serum virus neutralization assay is a serological test to detect the presence and functionality of the antibodies. It only detects antibodies that can block virus replication. Although neutralization assay is a highly sensitive and specific test, it only could be carried in a bio safety level 3 facility because of the handling of an active virus and it requires 5 to 7 days for the cell culture and virus amplification. It is for all these that could not be used as screening test.
Currently, there are many antibody detection methods on the market, ELISA, colloidal gold, or chemiluminescence [4,5]. Detection antibodies include IgG, IgM, IgA and total antibodies [6-9]. The antigen used for detection is mainly N proteins and S proteins of the virus. For the S proteins could be divided into full-length and fragments of receptor binding domain [4]. Since there is no unified standard, it is difficult to analyze and compare the test results. It is unclear which method we should use to reflect the real immunity to the virus [7,8].
To answer these questions, this study compared four commercially available kits, the viral N protein IgG and IgM ELISA kit, the viral S protein IgG and IgM ELISA kit; and a virus neutralization experiments.
MATERIALS AND METHODS
Recruitment of patients and specimen collection
Samples were collected from CDC. Totally there are 164 samples from positive Covid-19 patients confirmed by RT-PCR, male 94 (age range 1 to 69) and female 70 (age range 2 to 75). Covid-19 patient’s closely contacted groups have 184 samples, male 102 (age range 1 to 85) and female 82 (age range 10 months to 80). Health control group have 300 samples, male 269 (age 11-59) and female 31 (age range 21 to 62). All samples were leftover serums of other experiments.
COVID-19 S-IgG and S-IgM ELISA ASSAY kits were provided from Leide Biosciences Co., Ltd., (#2022-96 and #2023-96).
S protein was expressed in 293T cells and purified by Ni-sepharose. The purified S protein-based ELISA kits were used for the detection of IgG and IgM antibody against SARS-CoV-2 S protein. 5µl serum samples were diluted to 500µl and 100µl were used for test. According to the instruction of the kits, the cutoff value of S-IgG and S-IgM bothwere 0.3 of OD 450-630.
COVID-19 N-IgG and N-IgM ELISA ASSAY kits were provided by Zhongshan Biological Engineering Co., Ltd (# Y20200304). The recombinant nucleo capsid (rN) protein based ELISA kits were used for the detection of IgG and IgM antibody against SARS-CoV-2 N protein. According to the instruction of the kits, the cutoff value of N-IgM and N-IgG were 0.15 and 0.3 respectively. WhenA450-630 was below the cutoff value, the test was considered negative, and when A450-630 was greater than or equal to the cutoff value, the test was considered positive.
Vero-E6 cell line purchased from ATCC (CRL-1586)
SAS-COV2 virus come from Guangdong CDC (Gisaid library code is EPI_ISL_403934. Named as 20SF014/vero-E6/3. Plate reader iMark is from Bio-Rad.
Neutralization experiment
The SARS-COV-2 virus 20SF014/vero-E6/3 is isolated from a COVID-19 patient in Shenzhen in February. Stock virus was amplified by grown in Vero E6 cells. The virus stock was titrated by serial dilute by 10-fold. A confluent monolayer of Vero E6 cells was infected with 100 µl of each dilution in quadruplicate in 96-well plates. Cytopathic effect (CPE) was observed under microscope seven days after inoculation. The endpoint dilution leading to CPE in 50% of inoculated wells was designated as one 50% tissue culture infecting dose (TCID50).
Serial four-fold dilutions of heat-inactivated sera were made. The serum dilutions 240µl were mixed with equal volumes of 100 µl TCID 50 of SARS-CoV-2 as indicated. After 2 h of incubation at 37°C, 5% CO2 incubator, 100 μL of the virus–serum mixture was added in quadruplicate to Vero E6 cell mono layers in 96-well microtiter plates. Then, an additional 100 μL of culture medium IMM was added to each well and the plates incubated for 7 days at 37 °C in 5% CO2 in a humidified incubator. A serial 10-fold dilution of the virus: 100µl TCID50/50 μL, 10 µl TCID50/50 µl, 1 μL TCID50/50 μL and 0.1 TCID50/50 μL was made as control and loading into 8 wells of 96-well microtiter plates with additional 150 μL of culture medium MM. The CPE was read at 7 days post infection. The highest serum dilution that completely protected the cells from CPE in half of the wells was taken as the neutralizing antibody titer. More than 4 times dilution could protect the cells from virus infection was set as the cutoff of the positive. These procedures were carried in a biosafety level 3 facility.
ELISA binding assay
Each serum or plasma sample was tested at a dilution of 1:100 in sample dilution ELISA buffer (Supplied with each ELISA kits) and added to the ELISA wells of each plate for 1 h shaking with 250rpm at room temperature (or keep stand still at 37°C, follow the instruction of menu). After 5 times washing with washing buffer, horseradish peroxidase (HRP)-conjugated goat anti-human IgG or HRP-conjugated goat anti-human IgM were added for 30 minutes at shaking with 250rpm at room temperature (or keep stand still at 37°C, following the instructions). The ELISA plates were then washed five times with washing buffer. Subsequently, 100 μL of HRP substrate was added into each well. After 15 min incubation, the reaction was stopped by adding 50 μL of stop solution and read optical density (OD) within 15 minutes at 450nm and 630nm oni Mark microplate reader (BioRad).Comparing the sample OD value with the Reference Control’s OD value, it is positive when the S/CO is greater than 1.
STATISTICAL ANALYSIS
Correlations between serums ELISA OD, detection rates were assessed using Pearson’s correlation coefficients.
RESULT
- Performance comparison between four Covid-19antibody assay kits and one neutralization assay. Four ELISA kits, Anti-N IgG, Anti-N-IgM, Anti-S-IgG, Anti-S-IgM were used for testing. The Positive group was formed by 164 hospital-released Covid-19 patients. Covid-19 patients closely contacted group had 184 cases, and the control group had 300 cases. They were tested by these five assays for sensitivity and specificity. In the positive group, the detection rate of Anti-N IgG and neutralization experiments were the highest, both reaching 97.6%. In the closely contacted group, the detection rate of Anti-N IgG was 46.3%, the detection rate of neutralization experiment was 35.9%, and the detection rate of Anti-S-IgG was 34.7%. In the control group, the detection rate of Anti-N-IgG was 2.3% and the detection rate of Anti-N-IgM was 1%, but the detection rates of S-IgG, S-IgMwere zero (Table 1) (Supplementary Table 1).
|
Assays |
Positive Group(n=164) Positive rate (%) |
Closely contacted group (n=184) Positive rate (%) |
Negative control group(n=300) Positive rate (%) |
|
N-IgG(>0.5) |
160/164(97.6%) |
75/184(46.3%) |
2.3% |
|
N-IgM (>0.3) |
102/164(62.2%) |
47/184(25.5%) |
1% |
|
S-IgG (>0.3) |
153/164(93.9%) |
64/184(34.7%) |
0 |
|
S-IgM (>0.3) |
104/164(63.4%) |
37/184(20.1%) |
0 |
|
neutralization experiment |
160/164(97.6%) |
66/184(35.9%) |
0 |
Table 1: Four Covid-19 ELISA kits: N-IgG, N-IgM, S-IgG,S-IgM and neutralization experiment were tested with 164 positive patients’ group, 184 closely contacted group and 300 negative control group. Positive rates were compared as below.
- In the2X2 factorial design, the Anti-S-IgG ELISA and the neutralization assay were used to conduct a comparative study on 164 people that had closely contacted with covid-19patients. Their positive agreement rate was 59/64 (92.2%), negative agreement rate was 113/120 (94.2 %). The total agreement rate was 172/184 (93.4%) (Table 2).
|
|
Neutralization Assay |
|||
|
+ |
- |
Total |
||
|
S-IgG ELISA |
+ |
59 |
5 |
64 |
|
_ |
7 |
113 |
120 |
|
|
Total |
66 |
118 |
184 |
|
Positive agreement rate: 59/64=92.2%; Negative agreement rate: 113/120=94.2%; Positive predictive value, PPV=59/64=92.2%; Negative predictive value, NPV=113/120=94.2%; Total agreement rate: 172/184=93.4%
Table 2: 2X2 factorial design of neutralization assay and S-IgG assay in the Covid-19 closely contacted group.
- In the virus-neutralization assay, 164 samples of patients confirmed with Covid-19 and 184 samples of closely contacted person were tested. It was found that 160/164 (97.6%) of the confirmed patients did have virus-neutralizing capacity when discharged from the hospital. In the close contact group, 66/186 (35.9%) samples had virus neutralization activity (Figure 1) (Supplementary Table 2).
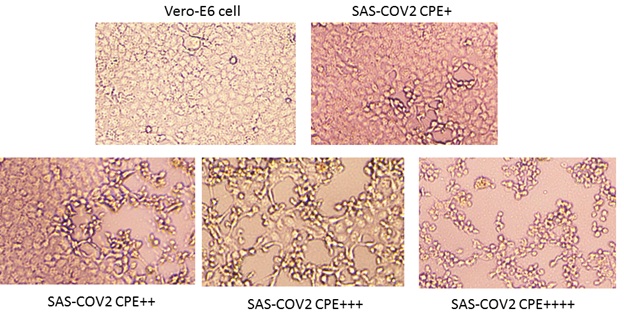 +: 25% cells was infected? ++: 50% cells was infected?+++?75% cells was infected?++++?100% cells were infected.
+: 25% cells was infected? ++: 50% cells was infected?+++?75% cells was infected?++++?100% cells were infected.
Figure 1: Neutralization experiment, SAS-COV2 virus infects Vero-E6 cells. There are four levels of Cytopathic effect (CPE). A confluent monolayer of Vero E6 cells was prepared in 96 well plate. Serial four-fold dilutions of heat-inactivated sera were made. The serum dilutions 240ul were mixed with equal volumes of 100 TCID50 of SARS-CoV-2 virus. After 2 h of incubation at 37°C, 5%CO2 incubator, 100μL of the virus–serum mixture were added in quadruplicate to Vero E6 cell monolayers in 96-well microtiter plates. Then the plates were incubated 7 days at 37°C in 5% CO2 in a humidified incubator. The CPE was read at 7 days post infection. The highest serum dilution that completely protected the cells from CPE in half of the wells was taken as the neutralizing antibody titre. More than 4 times dilution could protect the cells from virus infection was set as the cutoff of the positive. Below are 4 different levels of the virus infection.
- Covid-19 patients closely contacted group was separately into virus neutralization activity positive group (66 people) and negative group (118 people). The positive coincidence rate of N-IgG and S-IgG was identical, 59/66 (89.4%). But in the negative group, the detection rate of anti-N-IgG was16/118 (13.6%) where as anti-S-IgG was only 5/118 (4.2%). About 14/118(11.8%) people only produced antibodies against viral N protein (Figure 2).
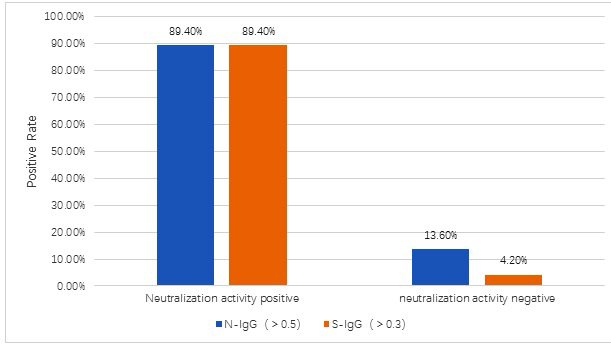
Figure 2: Comparing of N-IgG and S-IgG ELISA in the Covid-19 closely contacted group. Both assay had 59/66(89.4%) positive in the 66 neutralization positive samples. In the 118 persons which neutralization test negative group, N-IgG had 16positive (about 13.6%) and S-IgG had 5positive (about 4.2%). About 14 persons (11.8%) only produced antibody to the N protein of the SAS-COV2 virus.
DISCUSSION
The global epidemic of SARS-COV-2 caused the world's largest public health crisis with over 21 million infections and 766 thousand death till now. To answer the question which method could be used to detect the immunity after infection, 164 covid-19 patients, 184 high-risk individuals and 300 healthcare personnel were tested. The S- IgG assay showed 93.5% agreement with the virus neutralization experiment. It could be used to replace the high-risk, time-costly virus neutralization experiment. Normally, measuring neutralization antibodies requires the use of live virus, cells, highly skilled operators, and complex laboratory procedures that are generally less sensitive and require several days to obtain results. Conventional virus neutralization tests require live coronaviruses and a bio safety level 3 laboratory for processing. This report provided a simple method to test.
Results also showed that viral N-protein induced the highest antibody response. In the Covid-19 confirmed patient group, the closely contacted group and the health control group, the positive rate of N-IgG was the highest, 97.6%, 46.3%, and 2.3%, respectively; and the detection rate of S-IgG was slightly lower, 93.9%, 34.7% and 0 respectively. The positive rates of neutralization experiment were 97.6%, 35.9% and 0, respectively. Although Anti-N-IgG had high detection rate, the Anti-N-IgG and Anti-N-IgM in the control group’s detecting may be cross reactions owing to sequence the homology of the viral nucleoprotein, which need further confirm [10-16].
To answer the question whether each infection could produce protection immunity, a group of 184 people that had closely contacted with covid-19 patients were tested. Only 66 serum samples have neutralization activity. In those 118 persons that do not have neutralization activity, anti-N-IgG ELISA positive rate was14/118 (11.8%), which suggests that some infection did not produce protection antibodies. Some of the covid-19 patients only produced anti-N protein antibodies and no anti-S antibodies 5/164 (3.05%). Some of them neither had antibodies against S protein nor antibodies against N protein 5/164 (3.05%). How did these patients recover from the virus attack is still unknown? Whether they are vulnerable to virus re-infection is also unknown.
The United States plans to conduct antibody testing for nearly 10 million people [15]. If the N protein antigen is used, their test result will not reflect whether the patients have protective antibodies or not. If only the S protein antigen is used, some infected patients will be reported as false negative. A better way is to analyze both: IgG against S and IgG against N proteins. Thus, the highest infection rate and the effective immunity rate could be obtained simultaneously.
Molecular assays have been the gold standard to directly detect for the presence of viral genetic material in infected individuals. However, insufficient viral RNA at the point of detection may lead to false negative result.
FUNDING
This study was supported by grants from: The Key Research and Development Program of Guangdong Province (2019B111103001); Natural Science Foundation of Guangdong Province (2019A1515012121); The leading talents of Guangdong Province program (No.00201512); and the National Science and Technology Major Project (2018zx10732401). NIAID center of excellence for influnenza research and surveillance (HHSN2722014000006C).
AVAILABILITY OF DATA AND MATERIALS
The datasets used and/or analyzed in this study are available from the corresponding author upon reasonable request.
REFERENCES
- Syal K (2020) COVID-19: Herd immunity and convalescent plasma transfer therapy. J Med Virol.
- Lin CY, Xiang J, Yan MZ, Li H, Huang S, et al. (2020) Comparison of throat swabs and sputum specimens for viral nucleic acid detection in 52 cases of novel coronavirus (SARS-Cov-2) infected pneumonia (COVID-19). Clinical Chemistry and Laboratory Medicine (CCLM) 25: 1089-1094.
- Chen N, Zhou M, Dong X, Qu JM, Gong FY, et al. (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: descriptive study. Lancet 395: 507-513.
- Yuan M, Wu NC, Zhu X, Lee CD, So RTY, et al. (2020) A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV. Science 368: 630-633.
- Chan CM, Tse H, Wong SS, Woo PC, Lau SK, et al. (2009) Examination of seroprevalence of coronavirus HKU1 infection with S protein-based ELISA and neutralization assay against viral spike pseudotyped virus. J Clin Virol 45: 54-60.
- Okba NMA, Müller MA, Li W, Wang C, GeurtsvanKessel CH, et al. (2020) Severe acute respiratory syndrome coronavirus 2-specific antibody responses in coronavirus disease 2019 patients. Emerg Infect Dis 26: 1478-1488.
- Zhang B, Zhou X, Zhu C, Feng F, Qiu Y, et al. (2020) Immune phenotyping based on neutrophil-to-lymphocyte ratio and IgG predicts disease severity and outcome for patients with COVID-19. Front Mol Biosci 7: 57.
- Abbasi J (2020) The Promise and Peril of Antibody Testing for COVID-19. JAMA 323: 1881-1883.
- Lassaunière R, Frische A, Harboe ZB, Nielsen ACY, Fomsgard A, et al. (2020) Evaluation of nine commercial SARS-CoV-2 immunoassays. medRxiv.
- Yip MS, Leung JL, Li PH, Cheung CY, Dutry I, et al. (2016) Antibody-dependent enhancement of SARS coronavirus infection and its role in the pathogenesis of SARS. Hong Kong Med J 22: 25-31.
- Zhao J, Yuan Q, Wang H, Liu W, Liao X, et al. (2020) Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin Infect Dis.
- Wang KK, Tsang OTK, Leung WS, Tam AR, Wu TK, et al. (2020) Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet Infect Dis 20: 565-574.
- Liu W, Liu L, Kou G, Zheng Y, Ding Y, et al. (2020) Evaluation of Nucleocapsid and Spike Protein-based ELISAs for detecting antibodies against SARS-CoV-2. J Clin Microbiol 58: e00461-20.
- Bendavid E, Mulaney B, Sood N, Shah S, Ling E, et al. (2020) VID-19 Antibody Seroprevalence in Santa Clara County, California . medRxiv.
- An J, Liao X, Xiao T, Qian S, Yuan J, et al. (2020) Clinical characteristics of the recovered COVID-19 patients with re-detectable positive RNA test. medRxiv.
- Krammer F, Simon V (2020) Serology assays to manage COVID-19. Science 368: 1060-1061.
SUPPLEMENTARY TABLES
Supplementary Table 1:
Supplementary Table 2:
Citation: Liang L, Huang H, Zheng H, Zhang H, Zhou H, et al. (2020) S-IgG Assay is a Good Replacement of the High Risk, Time Costly Virus Neutralization Assay. J Clin Immunol Immunother 6: 042.
Copyright: © 2020 Lijun Liang, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.

