
To Explore the Mechanism of Polygoni Cuspidati Rhizoma et Radix-Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Pulmonary Nodules Complicated with Hyperlipidemia based on Network Pharmacology and Molecular Docking
*Corresponding Author(s):
Zhang YongshengDongfang Hospital Of Beijing University Of Chinese Medicine, Beijing, China
Tel:+86 13681298389,
Email:zhyshengcm@163.com
Chai Xinlou
College Of Traditional Chinese Medicine, Beijing University Of Traditional Chinese Medicine, Beijing, China
Tel:+86 15811484091,
Email:mmxin3@126.com
Abstract
Purpose: The purpose of this study is to explore the mechanism of Polygoni Cuspidati Rhizoma et Radix (PCRR) Salviae Miltiorrhizae Radix et Rhizoma (SMRR) in the treatment of Pulmonary Nodules (PN) with Hyperlipidemia (HLP) based on network pharmacology and molecular docking technology.
Methods: The active components and targets of PCRR-SMRR were screened by the TCMSP database. GeneCards, OMIM, and DisGeNET databases were used to collect targets related to PN and HLP. Upload drug targets and disease targets to the Venny platform for the intersection to find common targets between them. The Cytoscape 3.9 software and String database were used to construct the protein-protein interaction (PPI) network and obtain the core targets, and the targets were entered into the Metascape database for the Gene Ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. Finally, the top active ingredients are docked with core targets.
Results: A total of 75 active ingredients of PCRR-SMRR were screened, with 830 action targets. 3074 targets related to PN, and 1672 targets related to HLP were screened out. A total of 96 targets were obtained by taking the intersection of drug targets and disease targets to construct a PPI network, in which AKT1, TNF, IL6, IL1B, and TP53 were the core targets. GO enrichment analysis revealed molecular functions involving transcription factor binding, protein domain-specific binding, signaling receptor regulatory activity, etc. Biological processes concerning response to wounding, response to lipopolysaccharides, response to inorganic substances, blood vessel development, etc. KEGG enrichment analysis mainly includes the cancer pathway, AGE-RAGE pathway of signaling, HIF-1 signature pathway, P53 signaling pathway, NF-KB switching pathway, and JAK-STAT pathway, etc. The molecular docking results indicated that the core targets AKT1, TNF, IL6, IL1B, and TP53 displayed good binding ability with the active ingredients Przewaquinone E and beta-sitosterol with binding energy ≤ -5.0 Kcal/mol.
Conclusion: Based on network pharmacology and molecular docking techniques, this study reveals that the pharmacological effects of PCRR-SMRR on the treatment of PN combined with HLP are multi-target and multi-pathway modulated, and of course, further experiments are needed to verify the specific mechanisms subsequently.
Keywords
Hyperlipidemia; Molecular docking; Network pharmacology; Polygoni Cuspidati Rhizoma et Radix; Pulmonary nodules; Salviae Miltiorrhizae Radix et Rhizoma
Introduction
PN is round or irregularly shaped lesions less than or equal to 3 cm in diameter in the lungs, with imaging showing an increased density shadow, either single or multiple, with well-defined or indistinct borders [1]. With the popularity of Low-Dose Computed Tomography (LDCT), the detection rate of PN is increasing day by day, and their occurrence is closely related to air pollution, fume exposure, dietary and exercise habits, as well as endocrine, family history, and metabolic diseases. According to the results of the American Lung Cancer Screening Trial, the positive rate of lung nodules can be as high as 24.2%, and the proportion of malignant lung nodules can be as high as 3.6% [2], so early screening of lung nodules is also an important diagnostic method for lung cancer. HLP is a metabolic disorder in which the levels of lipids and lipoproteins in the blood are abnormally high [3]. A trial examining the phonetic characteristics of patients with PN showed that 31% had HLP [4]. Another study showed that dyslipidemia is a risk factor for malignant pulmonary nodules and that the occurrence of malignant pulmonary nodules is accompanied by abnormalities in fatty acid metabolism, including fatty acid synthesis, lipid mobilization, fatty acid beta-oxidation, and other metabolic processes [5]. PN have been clinically observed in association with HLP, but their specific mechanism is unknown, and appropriate pharmacological treatment is lacking. PCRR-SMRR are more often found in clinical experience formulas and have been used to good effect in the clinical treatment of PN or HLP by instructors. Previous experimental studies by our group have shown that PCRR-SMRR has protective effects on vascular endothelial cells. However, their specific pharmacological effects are not yet clear. Network pharmacology is a holistic approach to analyzing the molecular association between drugs and therapeutic targets from a systemic level and biological network perspective to reveal the systemic pharmacological mechanisms of drugs [6]. Therefore, this study investigated the possible targets and related pathways of PCRR-SMRR in treating PN combined with HLP by applying network pharmacology and validated by molecular docking technique, to provide an experimental basis for clinical application. The flowchart of this study is shown in figure 1.
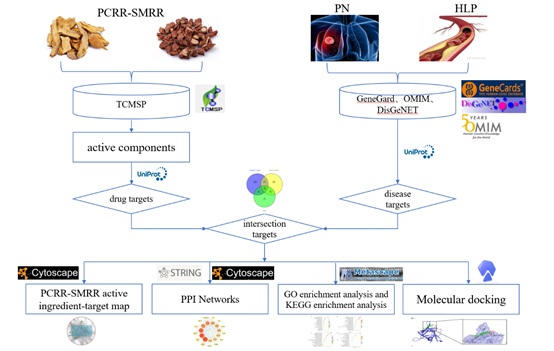 Figure 1: Detailed flowchart of the study design.
Figure 1: Detailed flowchart of the study design.
Methods
- Screening of active components and targets of PCRR-SMRR
Using PCRR-SMRR as keywords, we searched the TCMSP database [7] ( https://tcmsp-e.com/) separately to find their corresponding compounds, and set the composition screening conditions for oral bioavailability (OB) ≥ 30%, drug-like (DL) ≥ 0.18 [8], to obtain its effective active ingredients. The components of the resulting compounds were screened in the TCPSP database for target prediction. The Uniprot database [9] (http://www.uniprot.org/) was used to standardize the potential targets corresponding to the screened active components of PCRR-SMRR.
- Screening for disease targets in PN combined with HLP
Using PN and HLP as search terms, the GeneGard database [10] (https://www.genecards.org/), OMIM database [11] (https://www.omim.org), and DisGeNET database [12] (http://www.disgenet.org/) were searched separately. Combine and remove duplicates and false positives to obtain disease targets.
- Visualization of network regulation in the treatment of PN combined with HLP with PCRR-SMRR
The disease targets and drug targets were imported into Venny 2.1 software, and a Venn diagram was drawn to obtain their intersection targets. The data were imported into Cytoscape 3.9 software for visualization to obtain the active ingredient-target network diagram of Chinese medicine. The "rectangle" represents the target protein, the "hexagon" represents the compound components, and the "triangle" represents the common components of the two drugs [13].
- Constructing PPI networks and screening core targets
The screened drug-disease intersection targets were imported into the String 11.5 database [14] (http://string-db.org/), and the PPI network was constructed by selecting the biological species "Homo sapiens", setting the feasibility to > 0.4, hiding the free points, and leaving the rest as default settings [15]. The PPI network was imported into Cytoscape 3.9 software for visualization, and the CytoNCA was used for core target screening.
- GO Enrichment analysis and KEGG enrichment analysis
The intersecting targets obtained in step 1.3 were entered into the Metascape database [16] for Gene Ontology (GO) enrichment analysis and KEGG enrichment analysis, and then the data information was entered into the microbiology letter platform for visualization.
- Molecular docking
Molecular docking selected the five core targets screened in step 1.4 for validation against the three key components with high degree values. The protein crystal structures were downloaded from the PDB protein database [17] (http://www.pdb.org/), the relevant data were imported into AutodockTools 1.5.6 software for molecular docking, and the results of compound-protein docking were analyzed using PyMOL.
Results
- Screening and target prediction results of the active ingredients of PCRR-SMRR
The TCMSP database was searched for the two drugs and a total of 75 compound components were included, including 10 PCRR and 65 SMRR, as shown in table 1. The Uniprot database was used to search the above targets, standardize and unify the target gene names, and delete the related targets that could not be predicted. 830 gene targets were included, comprising 299 PCRR and 531 SMRR, and a total of 210 were obtained after deleting duplicate values.
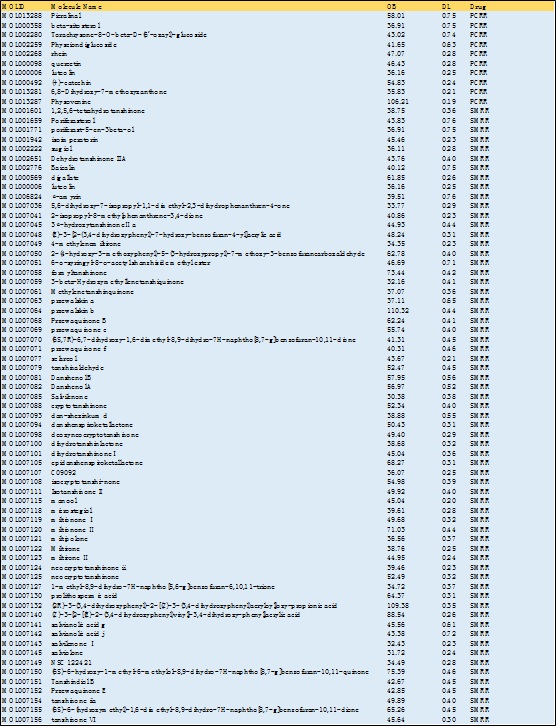 Table 1: PCRR-SMRR active ingredient information table.
Table 1: PCRR-SMRR active ingredient information table.
- PN combined with HLP disease target screening results
A total of 3074 targets related to PN and 1672 targets related to HLP were obtained by searching and screening disease targets in the GeneGard database, OMIM database, and DisGeNET database and combining and de-duplicating them.
- Network visualization analysis of PCRR-SMRR in the treatment of PN combined with HLP
The disease targets and 210 drug targets were imported into Venny 2.1 software to draw a Venn diagram, and a total of 96 intersection targets were obtained, as shown in figure 2. The visualization was performed using Cytoscape 3.9 software to obtain the relationship map between the herbal components and the target genes of PCRR-SMRR, and the results showed a total of 280 nodes and 899 edges. The larger the gene node display, the greater the number of connections to the drug component associated with the gene, see figure 3. Luteolin (hexagon A in the figure) is a common component of PCRR-SMRR. The results showed that quercetin was linked to the most genes (degree 141), indicating that it exerts the highest pharmacological effect, followed by przewaquinone E (degree 40), and beta-sitosterol (degree 38).
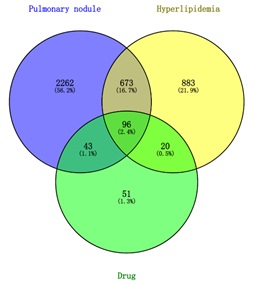 Figure 2: Venny diagram of drug targets and disease targets.
Figure 2: Venny diagram of drug targets and disease targets.
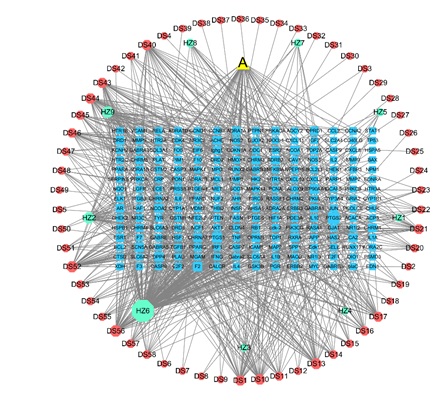 Figure 3: PCRR-SMRR active ingredient-target map; The hexagon in the figure shows the components of the herbal medicine, and the rectangle shows the target points, where the green part is the active component of PCRR and the orange part is the active component of SMRR; the blue part is the target point, and triangle A represents the common component of PCRR-SMRR.
Figure 3: PCRR-SMRR active ingredient-target map; The hexagon in the figure shows the components of the herbal medicine, and the rectangle shows the target points, where the green part is the active component of PCRR and the orange part is the active component of SMRR; the blue part is the target point, and triangle A represents the common component of PCRR-SMRR.
- PPI network construction and key target screening results
The 96 intersection targets were uploaded to String 11.5 database to obtain PPI network information. Also, save the TSV format file and upload it to Cytoscape 3.9 software to reconstruct the PPI network, see figure 4. The median of DC in the network is 44, the median of BC is 205.18799, and the median of CC is 0.66415437 obtained by topological analysis of the CytoNCA plug-in. With a median of greater than 3 parameters, the key targets AKT1 (degree 86), TNF (degree 85), IL6 (degree 83), IL1B (degree 76), and TP53 (degree 76) were finally obtained.
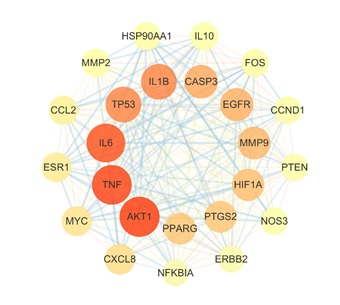 Figure 4: PPI Networks; The size and color depth of the nodes are adjusted according to the degree, the larger the node, the darker the color, indicating that the degree is larger, and the line between the nodes from thick to thin indicates that the number of edges from large to small.
Figure 4: PPI Networks; The size and color depth of the nodes are adjusted according to the degree, the larger the node, the darker the color, indicating that the degree is larger, and the line between the nodes from thick to thin indicates that the number of edges from large to small.
- Results of GO Enrichment Analysis and KEGG Enrichment analysis
Analysis was performed using the Metascape database to obtain cellular components, molecular functions, biological processes, and KEGG enrichment bubble maps. One of the cellular components, mainly related to the membrane raft, endoplasmic reticulum lumen, transcription regulator complex, vesicle lumen, etc., see figure 5a. Molecular functions, mainly related to transcription factor binding, protein domain specific binding, signaling receptor regulator activity, etc., see figure 5b. Biological processes, mainly seen in response to wounding, response to lipopolysaccharide, response to an inorganic substance, blood vessel development, etc., see figure 5c. KEGG enrichment analysis mainly includes the cancer pathway, AGE-RAGE pathway of signaling, HIF-1 signature pathway, P53 signaling pathway, NF-KB switching pathway, and JAK-STAT pathway, see figure 5d.
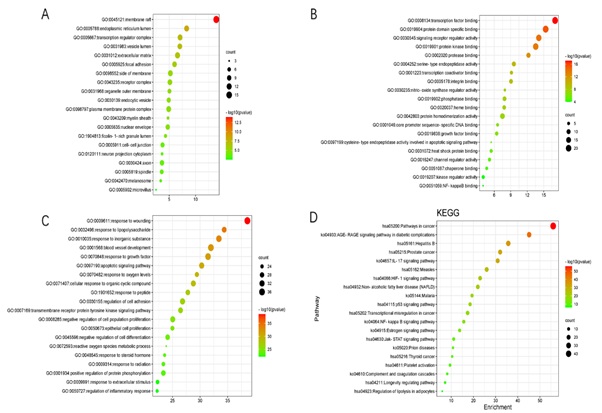 Figure 5: GO enrichment analysis and KEGG enrichment analysis; A: Cellular Components Map; B: Molecular functions Map; C: Biological Processes Map; D: KEGG Pathway Enrichment Map.
Figure 5: GO enrichment analysis and KEGG enrichment analysis; A: Cellular Components Map; B: Molecular functions Map; C: Biological Processes Map; D: KEGG Pathway Enrichment Map.
- Molecular docking results
The docking results of the top 5 key targets AKT1, TNF, IL6, IL1B, and TP53 with the key components quercetin, Przewaquinone E, and β-sitosterol are shown in table 2. beta-sitosterol had the lowest binding energy with 1L1B at -8.94kcal/mol, followed by beta-sitosterol with AKT1 at -7.83kcal/mol, beta-sitosterol with IL6 at -7.66 kcal/mol, Przewaquinone E with AKT1, -7.23 kcal/mol, and beta-sitosterol with TNF, -7.23 kcal/mol, as visualized in figure 6. The binding capacity of the key target to the active ingredient ≤ -5.0 Kcal/mol indicates good binding between the key target and the key ingredient, and the lower the binding energy, the more stable the docking. The results of this study showed that quercetin, progesterone E, and β-sitosterol, the core components of the PCRR-SMRR, dovetailed well with AKT1, TNF, IL6, IL1B, and TP53 molecules, the core targets of PN combined with HLP.
|
Key components |
Key targets |
||||
|
AKT1 |
TNF |
IL6 |
IL1B |
TP53 |
|
|
Quercetin |
-4.35 |
-4.43 |
-5.79 |
-5.96 |
-5.94 |
|
Przewaquinone E |
-7.23 |
-6.31 |
-6.83 |
-6.75 |
-6.4 |
|
beta-sitosterol |
-7.83 |
-7.23 |
-7.66 |
-8.94 |
-7.17 |
Table 2: Molecular docking results of key components and key targets (Kcal/mol).
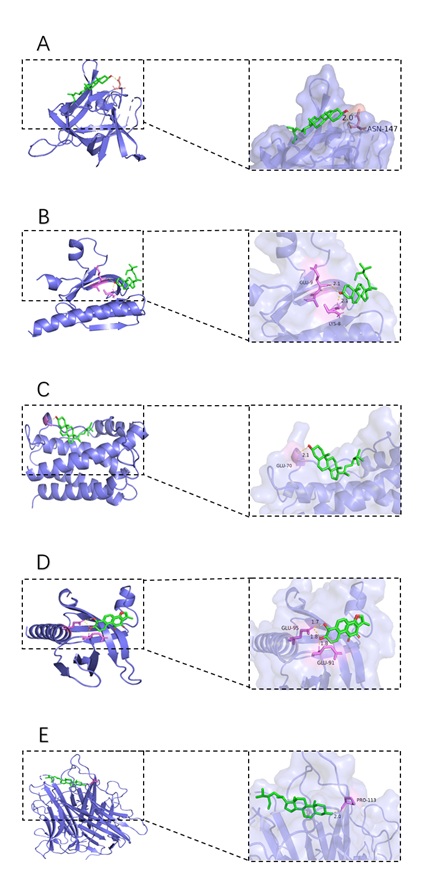 Figure 6: Schematic diagram of the docking of compounds with better binding activity to target protein molecules; A: beta-sitosterol and IL1B; B: beta-sitosterol and AKT1; C: beta-sitosterol and IL6; D: Przewaquinone E and AKT1; F: beta-sitosterol and TNF.
Figure 6: Schematic diagram of the docking of compounds with better binding activity to target protein molecules; A: beta-sitosterol and IL1B; B: beta-sitosterol and AKT1; C: beta-sitosterol and IL6; D: Przewaquinone E and AKT1; F: beta-sitosterol and TNF.
Discussion
According to Chinese medicine, PCRR is slightly bitter, cold in nature, and belongs to the damp-resolving drug. It has the function of clearing heat-toxin, dissipating blood stasis, and alleviating pain, relieving cough and phlegm. Modern pharmacology has shown that resveratrol glycosides, the main active ingredient of PCRR, have a calming and expectorant effect, and can also increase coronary blood flow and lower blood lipids [18]. SMRR is slightly bitter, slightly cold in nature, and belongs to the blood activating and stasis resolving drug with the effect of removing blood stasis for promoting tissue regeneration, cooling blood, and eliminating carbuncles. Modern pharmacology shows that Salvia has obvious anti-thrombotic, and anti-atherosclerotic, increase coronary flow, and improved microcirculation effects [19]. PN with HLP is the result of over-eating fatty, sweet, and thick flavors, which cause the body’s Qi, blood, and fluid to malfunction, resulting in phlegm, dampness, turbidity, stasis, and other pathological products. These pathological products infiltrate the veins and collaterals, resulting in damage to the lung collaterals and the formation of nodules over time. PCRR and SMRR, one on the side of affecting the qi, the other on the side of affecting blood circulation, the two drugs work together to regulate qi-flowing for activating stagnancy, promoting blood circulation for removing obstruction in collaterals. Therefore, it can play a proper therapeutic effect on this disease.
In this research, the TCMSP platform filtered out 75 active ingredients of the PCRR-SMRR, and Quercetin, Przewaquinone E, and β-sitosterol are the main pharmaceutical active ingredients that are strongly correlated with genetic targets, and luteolin, a common component of both. A total of 96 genes were intersected between the two drugs and PN combined with HLP, AKT1, TNF, IL6, IL1B, and TP53 are the key targets that have been screened. The primary pathways of action of PCRR-SMRR in the treatment of PN together with HLP were determined by GO and KEGG enrichment analysis, namely cancer pathway, AGE-RAGE pathway of signaling, HIF-1 signature pathway, P53 signaling pathway, NF-KB switching pathway, and JAK-STAT pathway. Among them, luteolin is a flavonoid compound.
Pro-inflammatory agents such as IL-1β, IL-6, and TNF-α can be suppressed by luteolin, and various signaling pathways such as NF-KB, JAK-STAT, and TLR signaling pathways can also be modulated [20]. There are three known isoforms of AKT, which are known to play an essential role in the management of protein composition, lipid metabolism, and cell proliferation. AKT1 activation is closely related to PI3K/AKT and VEGF pathways [21]. When AKT is activated by PI3K, it phosphorylates downstream substrates and the angiogenic capacity of ground glass nodules is upregulated, and angiogenesis is key to lung cancer progression [22]. These mechanisms are also involved in lipid metabolism, according to YE J’s research, insulin secretions after food intake result in the activation of the PI3K/AKT signaling pathway, and thus to an increase in glucose usage, greater lipid sedimentation in the body, and a reduced amount of free fatty acids in the circulating body [23].
Luteolin primarily targets PI3K / Akt pathway signals [24]. It protects endothelial cells and inhibits oxidative stress by removing ROS and regulating various cellular signaling mechanisms associated with ROS, thereby preventing the development of PN [25]. Quercetin regulation of lipid metabolism disorders is also associated with the AKT signaling pathway [26]. Xu D [27] stated that quercetin has an essential contribution in managing the progression of atherosclerosis induced by a high fructose diet, mainly by suppressing ROS and augmenting PI3K/AKT. Quercetin also regulates TP53 and thus acts as an anticancer agent. TP53 is the most tumor-associated oncogene, and studies have shown that mutations in genes including TP53 and EGFR drive the transformation of subsolid nodules to lung cancer [28]. According to Lee DH’s research [29], quercetin represses the build-up of HIF-1α through the inhibition of protein composition. The HIF-1α signaling pathway is a proven critical step in the metastatic advancement of tumors, which is implicated in tumor angiogenesis and aggressive migration [30]. The HIF-1 signaling pathway is participating in a number of occurrences such as secretion of pro-inflammatory factors, angiogenesis, regulation of vascular tone, leukocyte recruitment, and cell adhesion, which are postulated to be inflammation-related events that trigger PN and HLP [31]. Wu XJ [32] showed that quercetin regulates many signaling pathways through the whole body and overall, including the AGE-RAGE switching pathway in diabetic comorbidity. The AGE-RAGE signaling pathway is an essential pathway in glycolipid metabolism, and AGE/RAGE signaling has been characterized to promote vascular calcification by activating Nox-1 and by reducing SOD-1 expression to augment oxidative stress [33]. These binding targets form a signaling pathway that may account for the possible explanations of HLP in PN. β-sitosterol is a commonly occurring plant cholesterol component. Jayaraman S [34] revealed that β-sitosterol was shown to down-regulate the IKKβ / NF-κB and JNK signal pathways in fatty tissue and consequently suppress the production of the pro-inflammatory cytokines TNF-α and IL-6. IL6 is a pro-inflammatory cytokine that also has an important part to play in inflammation, atherosclerosis, and thrombosis. It can enhance the inflammatory response by upholding the exposure to adhesion molecules and can also impact the rate of lipid metabolism, particularly TC and TG metabolism. The research literature on Przewaquinone E is poorly reported, and its effective formation will be the direction of future research.
In summary, the active substances of PCRR and SMRR in the therapy of PN complicated by HLP are quercetin, przewaquinone E, β-sitosterol, and luteolin, of which luteolin is a common component of both. PCRR and SMRR mainly act on the targets AKT1, TNF, IL6, IL1B, and TP53, and may exert regulatory effects through the cancer pathway, AGE-RAGE pathway of signaling, HIF-1 signature pathway, P53 signaling pathway, NF-KB switching pathway, and JAK-STAT pathway. Of course, further experiments are needed to verify the specific mechanism.
Funding Statement
The Fifth National Training Program for Excellent Talents in Chinese Medicine by the State Administration of Traditional Chinese Medicine (State Office of Chinese Medicine Human Education Letter [2022] No. 1); The Sixth Batch of Academic Inheritance Training Program for Beijing Chinese Medicine Specialists (Beijing Chinese Medicine Section Word [2021] 169); The Third Zhongjing National Medical Seminar Training Program in Beijing (Beijing Chinese Medicine Science and Technology [2016] No. 114); National Science and Technology Major Project "Research and Development of New Chinese Medicines Based on Big Data" (2019ZX09201004-001-021); School-level project of Beijing University of Chinese Medicine.
References
- Gould MK, Donington J, Lynch WR, Mazzone PJ, Midthun DE, et al. (2013) Evaluation of individuals with pulmonary nodules: when is it lung cancer? Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 143: 93-120.
- Wu H, Wang Q, Liu Q, Zhang Q, Huang Q, et al. (2020) The Serum Tumor Markers in Combination for Clinical Diagnosis of Lung Cancer. Clin Lab 66.
- Stone NJ, Robinson JG, Lichtenstein AH, Merz CNB, Blum CB, et al. (2014) 2013 ACC/AHA guideline on the treatment of blood cholesterol to reduce atherosclerotic cardiovascular risk in adults: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J Am Coll Cardiol 63: 2889-2934.
- Song XY, Wang SJ, Xu ZX, Hao YM, Feng L, et al. (2020) Preliminary study on phonetic characteristics of patients with pulmonary nodules. Journal of integrative medicine 18: 499-504.
- Dou T (2021) Metabolomics-based targeting of free fatty acid profiles for screening potential biomarkers of malignant lung nodules. Dalian Medical University, China.
- Li S (2021) A guide to network pharmacology evaluation methods. World Journal of Traditional Chinese Medicine 16: 527-532.
- Ru J, Li P, Wang J, Zhou W, Li B, et al. (2014) TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform 6: 13.
- Huang SJ, Mu F, Li F, Wang WJ, Zhang W, et al. (2020) Systematic Elucidation of the Potential Mechanism of Erzhi Pill against Drug-Induced Liver Injury via Network Pharmacology Approach. Evid Based Complement Alternat Med 2020: 6219432.
- UniProt Consortium (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 49: 480-489.
- Stelzer G, Dalah I, Stein TI, Satanower Y, Rosen N, et al. (2011) In-silico human genomics with GeneCards. Hum Genomics 5: 709-717.
- Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A (2015) OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res 43: 789-798.
- Piñero J, Anguita JMR, Pitarch JS, Ronzano F, Centeno E, et al. (2020) The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res 48: 845-855.
- Xinxin P, Qing Z, Zining P, Yage Z, Xiujie S, et al. (2022) Exploring the mechanism of hirudin in the treatment of diabetic kidney disease using network pharmacology combined with molecular docking verification. J Tradit Chin Med 42: 586-594.
- Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, et al. (2021) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49: 605-612.
- Hong J, Ding J, Hong HH, Xu XW, Pan B, et al. (2022) Identifying the Mechanism of Polygoni Cuspidati Rhizoma et Radix in Treating Acute Liver Failure Based on Network Pharmacology and Molecular Docking. Gastroenterol Res Pract 2022: 2021066.
- Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, et al. (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10: 1523.
- Noguchi T, Akiyama Y (2003) PDB-REPRDB: a database of representative protein chains from the Protein Data Bank (PDB) in 2003. Nucleic Acids Res 31: 492-493.
- Fan HT, Ding SL, Lin HS (20132) Pharmacological of Polygoni cuspidati rhizoma. Zhongguo Zhong Yao Za Zhi 38: 2545-2548.
- Shan XX, Hong BZ, Liu J, Wang GK, Chen WD, et al. (2021) Review of chemical composition, pharmacological effects, and clinical application of Salviae Miltiorrhizae Radix et Rhizoma and prediction of its Q-markers. Zhongguo Zhong Yao Za Zhi 46: 5496-5511.
- Gendrisch F, Esser PR, Schempp CM, Wölfle U (2021) Luteolin as a modulator of skin aging and inflammation. Biofactors 47: 170-180.
- Fan X, Lin L, Wang J, Wang Y, Feng A, et al. (2018) Genome profile in a extremely rare case of pulmonary sclerosing pneumocytoma presenting with diffusely-scattered nodules in the right lung. Cancer biology & therapy 19: 13-19.
- Lu T, Yang X, Shi Y, Zhao M, Bi G, et al. (2020) Single-cell transcriptome atlas of lung adenocarcinoma featured with ground glass nodules. Cell discovery 6: 69.
- Ye J, Li L, Hu Z (2021) Exploring the Molecular Mechanism of Action of Yinchen Wuling Powder for the Treatment of Hyperlipidemia, Using Network Pharmacology, Molecular Docking, and Molecular Dynamics Simulation. BioMed research international 2021: 9965906.
- Bagli E, Stefaniotou M, Morbidelli L, Ziche M, Psillas K, et al. (2004) Luteolin inhibits vascular endothelial growth factor-induced angiogenesis; inhibition of endothelial cell survival and proliferation by targeting phosphatidylinositol 3'-kinase activity. Cancer Res 64: 7936-7946.
- Imran M, Rauf A, Izneid TA, Nadeem M, Shariati MA, et al. (2019) Luteolin, a flavonoid, as an anticancer agent: A review. Biomed Pharmacother 112: 108612.
- Peng J, Li Q, Li K, Zhu L, Lin X, et al. (2017) Quercetin Improves Glucose and Lipid Metabolism of Diabetic Rats: Involvement of Akt Signaling and SIRT1. J Diabetes Res 2017: 3417306.
- Xu D, Hu MJ, Wang YQ, Cui YL (2019) Antioxidant Activities of Quercetin and Its Complexes for Medicinal Application. Molecules 24: 1123.
- Li Y, Li X, Li H, Zhao Y, Liu Z, et al. (2020) Genomic characterisation of pulmonary subsolid nodules: mutational landscape and radiological features. The European respiratory journal 55: 1901409.
- Lee DH, Lee YJ (2008) Quercetin suppresses hypoxia-induced accumulation of hypoxia-inducible factor-1alpha (HIF-1alpha) through inhibiting protein synthesis. J Cell Biochem 105: 546-553.
- Cai WB, Zhang Y, Cheng R, Wang Z, Fang SH, et al. (2012) Dual inhibition of plasminogen kringle 5 on angiogenesis and chemotaxis suppresses tumor metastasis by targeting HIF-1α pathway. PLoS One 7: 53152.
- Yuan G, Shi S, Jia Q, Shi J, Shi S, et al. (2021) Use of Network Pharmacology to Explore the Mechanism of Gegen (Puerariae lobatae Radix) in the Treatment of Type 2 Diabetes Mellitus Associated with Hyperlipidemia. Evid Based Complement Alternat Med 2021: 6633402.
- Wu XJ, Zhou XB, Chen C, Mao W (2016) Systematic Investigation of Quercetin for Treating Cardiovascular Disease Based on Network Pharmacology. Comb Chem High Throughput Screen 22: 411-420.
- Kay AM, Simpson CL, Stewart JA (2016) The Role of AGE/RAGE Signaling in Diabetes-Mediated Vascular Calcification. J Diabetes Res 2016: 6809703.
- Jayaraman S, Devarajan N, Rajagopal P, Babu S, Ganesan SK, et al. (2021) β-Sitosterol Circumvents Obesity Induced Inflammation and Insulin Resistance by down-Regulating IKKβ/NF-κB and JNK Signaling Pathway in Adipocytes of Type 2 Diabetic Rats. Molecules 26: 2101.
Citation: Rui L, Yongsheng Z, Xinlou C, Tianshu Y, Haotian X, et al. (2023) To Explore the Mechanism of Polygoni Cuspidati Rhizoma et Radix-Salviae Miltiorrhizae Radix et Rhizoma in the Treatment of Pulmonary Nodules Complicated with Hyperlipidemia based on Network Pharmacology and Molecular Docking. J Altern Complement Integr Med 9: 340.
Copyright: © 2023 Li Rui, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.

