
Meals, Preparation Environment and Hands of Food Handlers - Microbiological Status in Hospital Kitchens
*Corresponding Author(s):
Julia Arantes GalvãoDepartment Of Veterinary Medicine, Federal University Of Paraná, Curitiba, Brazil
Tel:+55 4192255697,
Email:juliaarantesgalvao@gmail.com
Abstract
The aim of this study was to assess the microbiological conditions of ready-to-serve meals, the preparation environment and the hands of food handlers at two public hospital allocated in São Paulo State, Brazil. Were evaluated 480 samples of food, equipment, utensils, drains and hands of the staff from three kitchen sectors of two public hospitals in Brazil. Total coliforms were found in 116 (24.17%) samples, Escherichia coli in nine (2.08%) samples and coagulase-positive Staphylococcus in 17 (3.54%) samples, two of these strains showed gene encoders for classic entero toxins production. Coagulase-Negative Staphylococcus (CNS) occurred in 98 (20.41%) samples, in which 19 gene encoders for classic enterotoxins production were detect, and Listeria monocytogenes occurred in four (0.83%) samples. Salmonella were not detect. The microbiological quality of most samples evaluated was considered satisfactory; however, the presence of L. monocytogenes and other microorganisms, even at low frequency and with low counts, represents a risk of cross contamination of the food items, which can transmit pathogens to the patients, as well as forming bio film. The great concern is that Listeria and CNS were not included at sanitary micro biological standards for foods in Brazil.
Keywords
INTRODUCTION
Enteric pathogens are harmless to most of the population, never the less they may cause illness and even death in susceptible individuals, particularly those immune compromised. A total of 1022 outbreaks of no socomial infections in the United States, United Kingdom, France, Canada, Germany, the Netherlands and Spain were report from 1966 to 2002 and only 3.32% of these were identified as food borne diseases [1].
However, the data in the literature do not reveal the true incidence of diseases originated from food in hospital units since most cases are not reported. The main factors that contribute to the occurrence of food borne diseases are poor personal hygiene habits of the food handlers, the cooking and storage of food at inappropriate temperatures, the acquisition of raw materials from unreliable sources and the use of poorly sanitized equipment. Since patients have a greater risk of becoming ill when exposed to potential food borne pathogens and given that the food services need to provide a wide variety of foods, it is essential that appropriate food handling practices are maintained [2].
The aim of this study was to evaluate the hygiene-sanitary quality of food prepared in two kitchens at public hospitals in São Paulo State, Brazil, as well as to study the dynamics of the contamination of ready-to-serve meals from utensils, equipment, the environment and the food handlers involved in the process.
MATERIAL AND METHODS
• Milk dispensary: an isolated area, with restricted access to staff using special clothing and hand sanitation. Utensils and equipment were restricted to this environment;
• General kitchen: the patients without dietary restrictions, staff and students had the meals prepared here. At HA the equipment and utensils were of exclusive use at this area, while at HB it has shared with the Special Diet Kitchen. The salads served at HA are acquired ready-to-serve, only being divided into portions in this environment, while at HB, the salads are sanitized, prepared and divided into portions in this environment; and
• Special Diet Kitchen, where the meals were prepared for patients with some dietary restriction - diabetic, hypertensive, etc.
Around 100 ml/g of each food product (meat, rice, soup, beans, chicken meat, potato, and other cooked food available for patients) were collected. For the equipment and utensils a smear of the surface delimited by molds, carried out with swabs, which were transfer to tubes containing 10 ml of Letheen broth. It was collected smears from the drains using sterile sponges soaked in 10 ml of Letheen broth and transferred to Whirl-Pak® bags containing 100 ml of Letheen broth.
The staff hands area was measured as shown in figure 1, so rinsed for around 1 min. in Whirl-Pak® bags containing 100 ml of Letheen broth. At each visit at the Milk Dispensary, was collected a sample of milk or substitute at feeding bottle, two swabs of utensils, two swabs of equipment and two rinses of the hands of the staff. At the General Kitchen a hot meal, a cold meal, two swabs of utensils, two swabs of equipment, two rinses of the hands of the staff and a swab of a drain was collected; and in the Special Diet Kitchen the same protocol was designed, except for the cold meal (that was the same). At HB the General and the Special Diet Kitchens shared the utensils and equipment, so the total swabs were sampled from the same place. The samples were stored in isothermal plastic boxes containing recyclable ice and transported to the laboratory where it was analyze on the same day as the collection.
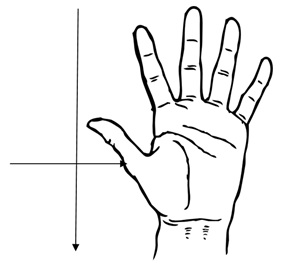 Figure 1: Measure the area of the hands of food handlers (cm²).
Figure 1: Measure the area of the hands of food handlers (cm²).Microbiological analysis
To test for Salmonella spp. variable volumes of 1% buffered peptone water was added to test bags or tubes, observing a ratio of 1:9 (sample/diluent). The same procedure was used to test for L. monocytogenes, in this case, employing Listeria broth enrichment, and for the enumeration of coagulase-positive and negative Staphylococcus, total coliforms and E. coli tenfold dilution were carried out in 0.85% saline solution.
All culture media were of the brand Difco®, with the exception of PetrifilmTM EC (3M®) and the agar Cromo Cen Listeria (base Agar Listeria Ottaviani Agosti; Biocen®).
Complementary analysis
| Gene | Primer | Sequence | Base pairs | Annealing Temperature |
| sea[7] | SEA-1SEA-2 | ttggaaacggttaaaacgaagaaccttcccatcaaaaaca | 120 | 50°C |
| seb[7] | SEB-1SEB-2 | tcgcatcaaactgacaaacggcaggtactctataagtgcc | 478 | |
| sec[7] | SEC-1SEC-2 | gacataaaagctaggaatttaaatcggattaacattatcc | 257 | |
| sed[7] | SED-1SED-2 | ctagtttggtaatatctccttaatgctatatcttataggg | 137 | |
| see[8] | SEE-1SEE-2 | aggttttttcacaggtcatcccttttttttcttcggtcaatc | 209 |
For the amplification, the Gene Amp PCR System 9700 (Applied Bio systems®) was used, with the following program: 94ºC/7 min (initial denaturation), followed by 30 cycles of 94ºC/30s, 50ºC /30s and 72ºC/30s, with a reduction of 1ºC per cycle in the annealing phase until reaching 45ºC. For the final period, 72°C had applied for 5 min. In all of the reactions the strains ATCC 13565 (sea), ATCC 14458 (seb), ATCC 19095 (sec), FRI 361 (sed), and ATCC 27664 (see) were used as positive controls and ultrapure water free of nucleases was used as the negative control. Universal primers originating from 16S rRNA forming a product of 371bp has used as the internal control [9].
The products of the PCR reactions were submitted to electrophoresis (Electrophoresis Power Supply Model EPD 600 - Amersham-Pharmacia Biotech Inc.) in 1.5% agarose gel (Prodinasa®) in Tris-boric acid-EDTA 1X (TBE 1X) buffer and developed with 1 µL of SYBR® safe (Invitrogen®)/10 ml of agarose gel. Comparative analysis were carried out with a label of 50 bp (LGC Biotecnologia®), and photographs of the DNA fragments has taken with an image analyzer (Alphaimager - AlphaEase FC Software - AlphaInotech Corporation®). The amplification of the internal control verified the good performance of the PCR and the absence of inhibitory agents in the reaction and extraction.
The L. monocytogenes strains were tested in API Listeria® (Biomérieux®) and serotyping [10], through pulsed-field gel electrophoresis (PulseNet protocol) at the Pharmaceutical Sciences Department of University of São Paulo (Faculdade de Ciências Farmacêuticas da Universidade de São Paulo).
All the results were interpreted according to the limits established by Brazilian legislation (Table 2). The foods contaminated by L. monocytogenes were classified as unsafe products.
| Food Group | Recommended Tolerance for Sample | |||
| Total Coliforms | Coliforms 45ºC | Coagulase-Positive Staphylococcus | Salmonella | |
| Ready-to-serve or instant products which will be consumed by children over one year of age after the addition of liquids | 20 | 1 | 50 | Absent |
| Ready-to-serve or instant products which will be consumed by babies under one year of age after the addition of liquids | 10 | Absent | Absent | Absent |
| Infant formulas for premature babies | 10 | Absent | Absent | Absent |
| Bottled water for the preparation of feeding bottles | Absent | - | - | - |
| Pasteurized milk | - | 4 | - | Absent |
| Fresh unprocessed vegetables prepared for consumption | - | 100 | - | Absent |
| Ready-to-serve meals (ready-to-serve foods of kitchens, restaurants, etc.) based on cooked meat, fish, eggs and so on. | - | 20 | 1000 | Absent |
Ethical aspects
RESULTS AND DISCUSSION
Total coliforms were found in 116 (24.17%) samples. At HA 52.58% of the samples were contaminated, the Milk Dispensary showed 13.79% of the samples at the range 1.0×10° to 8.8×10³ CFU/ml or cm². A feeding bottle was inappropriate [11] for consumption by premature babies or by children both under and over one year of age, since it represents an infant food, which contains 1.3×10² CFU/ml of total coliforms. At General Kitchen 19.83% of the samples showed 1.0×10¹ to 1.1×104 CFU/g or cm², and from the Special Diet Kitchen 18.97% of the samples showed 7.0×10° to 7.0×10³ CFU/g or cm². At HB Milk Dispensary 8.62% of the samples were contaminated with 2.0×10° to 2.0×10³ CFU/ml or cm²,from General Kitchen, 11.21% showed <1.0×10¹ to 2.3×10³ CFU/g or cm², from the Special Diet Kitchen, 7.76% of the samples showed 1.0×10¹ to 9.2×10³ CFU/g, and from the utensils and equipment used both in the General and Special Diet Kitchens, 19.83% showed 1.0×10° to 1.7×104 CFU/cm².
Nine (2.08%) samples were contaminated by E. coli, and none of these was originated from HA or from the Milk Dispensary of HB. At HA, the general kitchen had 33.33% of the samples with counts by <1.0×10¹ to 4.0×10¹ CFU/g, the Special Diet Kitchen 22.22% with 1.1×10¹ to 2.0×10¹ CFU/g, one of that, a hot meal was inappropriate for consumption, since it was contaminated with 2.0×10¹ CFU/g of E. coli[11].Furthermore, 44.44% from the utensils and equipment used in the General and Special Diet Kitchens at HB had 2.0×10° to 2.5×10¹ CFU/cm².
The results obtained for the total coliforms and E. coli counts of the samples collected from the hands of the food handlers reveal that good hygiene practices were adopting in relation to the hands at both units. In another study, of 180 samples analyzed, 8% was contaminate with E. coli [12].
Coagulase-positive Staphylococcus occurred in 17 (3.54%) samples, two of these strains showed gene encoders for classic enterotoxins production, 12.5% of these samples came from hands of food handlers at the Milk Dispensary (1.0×10¹ to 3.0×10² CFU/cm²), 18.75% from meals and hands of food handlers at the General Kitchen (1.0×10¹ to 1.0×10² CFU/g or cm²) and 18.75% from the Special Diet Kitchen (9.5×10¹ to 1.1×10² CFU/g or cm²).
At HB the samples contaminated by coagulase-positive Staphylococcus was 12.5% originated from the Milk Dispensary (it was a hand of food handler with 1.0×10² CFU/cm²), 12.5% from the General Kitchen (<1.0×10² to 1.0×10² CFU/g or cm²), 18.75% from the Special Diet Kitchen (4.3×10¹ to 2.0×10² CFU/g or cm²) and 6.25% from the utensils and equipment used in the General and Special Diet Kitchens (an equipment with 4.3×10³ CFU/cm²).
The strains with gene encoders for production of classic enterotoxins has detected at a blender (sea e sec) from General Kitchen of HA and, at the hands of a food handler (seb e sec) from General Kitchen of HB.
Regarding the presence of coagulase-positive Staphylococcus, a low frequency of this microorganism was found and when it was present, it did not reach a significant count, considering national legislation [11] and the number of viable cells required for the production of toxins, which is over 105 CFU/g of food [13].
Other authors, who evaluated 70 samples of salads to be served to hospitalized individuals in Turkey, have obtain results of greater concern, eight (11%) of the samples being contaminated by coagulase-positive Staphylococcus, with counts ranging from 1.0×10³ to 1.0×104 CFU/g12.Coagulase-Negative Staphylococcus (CNS) occurred in 98 (20.41%) samples, in which 19 gene encoders for classic enterotoxins production were detect (Table 3).
| Hospital A | Toxin gen | Milk Dispensary | General Kitchen | Special Diet Kitchen | |||
| Sample | Strain | Sample | Strain | Sample | Strain | ||
| sea | - | - | - | - | - | - | |
| seb | - | - | - | - | - | - | |
| sec | - | - | 1 Hand | S. warnerri | - | - | |
| sed | - | - | - | - | - | - | |
| see | 1 Hand | S. warnerri | 1 Hand | S. warneri | 1 Food handler / 2 Hands | S. epidermidis | |
| - | - | 3 Food handler / 4 Hands | S. saprophyticus | 3 Food handler / 3 Hands | S. warneri | ||
| - | - | 1 Tomato sauce | S. saprophyticus | - | - | ||
| Hospital B | sea | - | - | - | - | - | - |
| seb | - | - | 1 Spoon | S. xylosus | - | - | |
| - | - | 1 Fork | S. xylosus | - | - | ||
| - | - | 1 Food handler / 1 Hand | S. saprophyticus | - | - | ||
| sec | 1 Food handler / 1 Hand | S. lugdunensis | - | - | 1 Food handler / 1 Hand | S. warneri | |
| - | - | - | - | 1 Fork | S. xylosus | ||
| sed | - | - | - | - | - | - | |
| see | - | - | 1 Food handler/1 Hand | S. caprae | - | - | |
Table 3: Coagulase-negative Staphylococcus strains from Hospitals A and B with gene encoders for production of classic enterotoxins.
At HA 14.17% of the contaminated samples with CNS came from the Milk Dispensary, none of that was by feeding bottles and the count was by 3.1×10¹ to 1.3×104 CFU/cm². The General Kitchen had 18.9% of the contaminated samples (1.8×10¹ to 1.6×105 CFU/g or cm²), and the Special Diet Kitchen18.11% of the samples (5.1×10¹ to 6.8×104 CFU/g or cm²), totalizing 51.18% of the samples contaminated by coagulase-negative Staphylococcus. At HB 14.96% of the contaminated samples had been originated from the Milk Dispensary, in the same way observed in HA none of that was by feeding bottles, and the counts were by 1.1×10¹ to 1.4×104 CFU/cm². From the General Kitchen were 11.02% of the samples (<1.0×10² to 4.1×104 CFU/g or cm²), 11.81% from the Special Diet Kitchen (3.1×10¹ to 5.0×104 CFU/g or cm²), and 11.02% from the utensils and equipment used in the General and Special Diet Kitchens (3.3×10¹ to 1.4×104 CFU/g or cm²).
The high counts of coagulase-negative Staphylococcus should not been disregarded, since these micro organisms are potential producers of SE. It has created a great concern, especially because this pathogen was not included at national legislation [11] for the hospital food, hands of staff or kitchens environment, which hinders the implementation of corrective/preventative actions. However, the production of toxins has not tested in this study.
Listeria monocytogenes occurred in four (0.83%) samples. It was detected at a drain from HA (serotype 1/2b, 3b, 7), and at two samples of drains (serotype 4a, 4c and 1/2b, 3b, 7) and at an equipment (a blender-serotype not identified) at HB. In addiction an equipment (a vase) from HA and a drain from HB were contaminated with L. innocua.
There were none ready-to-serve meal contaminated by L. monocytogenes. In another study, 29 of 950 sandwiches prepared in a hospital in the United Kingdom were contaminate by L. monocytogenes, and for one sample, the count was 1.2×10³ CFU/g [14].
The detection of Listeria at the equipment and environment created a great concern, especially because the absence of this pathogen was not included at national legislation (Table 2) for the hospital kitchens environment. An improvement in relation to the sanitation and disinfection of the equipment and drains was poignantly recommend, since the presence of L. monocytogenes and yours marker (L. innocua) is unacceptable particularly at an environment that prepares meals for hospitalized individuals.
There was none sample contaminated by Salmonella spp. The absence of pathogens in the final meals, as well as the low counts for E. coli, detected only in the cold meals and within the standards established by legislation [11], lead to the conclusion that the units present good hygiene-sanitary control in relation to their operations.
So, the microbiological quality of most samples evaluated has considered satisfactory; however, the presence of L. monocytogenes and other microorganisms, even at low frequency and with low counts, represents a risk of cross contamination of the food items, which can transmit pathogens to the patients, as well as the possible formation of a biofilm.
The detection of Listeria at the equipment and environment created a great concern, especially because this pathogen was not included at national legislation (Table 2) for the hospital kitchens environment.
An improvement in relation to the sanitation and disinfection of the equipment and drains was poignantly recommend, since the presence of L. monocytogenes and yours marker (L. innocua) is unacceptable particularly at an environment that prepares meals for hospitalized individuals.
ACKNOWLEDGMENT
We thank for Maria Teresa Destro (Universidade de São Paulo, Faculdade de Ciências Farmacêuticas, Departamento de Alimentos e Nutrição Experimental) for confirmation and serotyping of Listeria strains. We also thank the nutritionists and the hospital management for permit this study.
REFERENCES
- Gastmeier P, Stamm-Balderjahn S, Hansen S, Nitzschke-Tiemann F, Zuschneid I, et al. (2005) How outbreaks can contribute to prevention of nosocomial infection: Analysis of 1022 outbreaks. Infect Control Hosp Epidemiol 26: 357-361.
- Réglier-Poupet H, Paraina C, Beauvais R, Descamps P, Gillet H, et al. (2005) Evaluation of the quality of hospital food from the kitchen to the patient. J Hosp Infect 59: 131-137.
- Andrews WH, June GA, Sherrod PS, Hammack TS. Amaguana, et al. (1998) Food and Drug Administration. Bacteriological Analytical Manual, Gaithersburg: AOAC International, Maryland, USA.
- Pagotto F, Daley E, Farber JM (2011) Enumeration of Listeria monocytogenes in food. Compendium of analytical methods, MFLP-74. Health Canada, Ontario, Canada.
- Ministry of Agriculture, Livestock and Supply (2003) Normative Instruction Nº 62. Methods for microbiological analysis of food of animal origin and water. Brasília, Brazil.
- Cunha ML, Sinzato YK, Silveira LV (2004) Comparison of Methods for the Identification of Coagulase negative Staphylococci. MemInst Oswaldo Cruz 99: 855-860.
- Johnson WM, Tyler SD, Ewan EP, Ashton FE, Pollard DR, et al. (1991) Detection of genes for enterotoxins, exfoliative toxins, and toxic shock syndrome toxin1 in Staphylococcus aureus by the polymerase chain reaction. J Clin Microbiol 29: 426-430.
- Mehrotra M, Wang G, Johnson WM (2000) Multiplex PCR for detection of genes for Staphylococcusaureus enterotoxins, exfoliative toxins, toxic shock syndrome toxin 1, and methicillin resistance. J Clin Microbiol 38: 1032-1035.
- Greisen K, Loeffelholz M, Purohit A, Leong D (1994) PCR primers and probes for the 16S rRNA gene of most species of pathogenic bacteria, including bacteria found in cerebrospinal fluid. J Clin Microbiol 32: 335-351.
- Doumith M, Buchrieser C, Glaser P, Jacquet C, Martin P (2004) Differentiation of the Major Listeria monocytogenes Serovars by Multiplex PCR. J Clin Microbiol 42: 3819-3822.
- Ministry of Health Resolution (2001) RDC Nº 12. Technical Regulation on microbiological standards for foods, Brazil.
- Ayçiçek H, Sarimehmetoglu B, Çakiroglu S (2004) Assessment of microbiological quality of meals sampled at the meal serving units of a military hospital in Ankara, Turkey. Food Control 15: 379-384.
- Tranter HS (1990) Foodborne Staphylococcal illness. Lancet 336: 1044-1046.
- Meldrum RJ, Smith RM (2007) Occurrence of Listeria monocytogenes in sandwiches available to hospital patients in Wales, United Kingdom. J Food Protect 70: 1958-1960.
Citation: Galvão JA, d’Ovidio L, Buzi KA, Yamatogi RS, Rall VLM, et al. (2017) Meals, Preparation Environment and Hands of Food Handlers - Microbiological Status in Hospital Kitchens. J Food Sci Nut 3: 018.
Copyright: © 2017 Julia Arantes Galvão, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.

