
The Karyotype of the Forensically Important Flesh Flies (Diptera: Sarcophagidae), Mini Review
*Corresponding Author(s):
Ghada Mohamed El-BassionyDepartment Of Entomology, Faculty Of Science, Cairo University, Cairo, Egypt
Tel:+201144002298,
Email:ghada@sci.cu.edu.eg
Abstract
Maggots of flesh flies were known as one of the main players in depletion of the earthily decomposed remains. Sarcophagidae colonized the carrion few minutes after death, and were known as the initial colonizers in several research papers. In consequence, they could be used in the minimum Post-Mortem Intervals (PMImini) estimation. Also, some sarcophagid species proved their capability in detecting toxins/ drugs in carcasses in the entomotoxicology field. The difficulty to identify sarcophagid females and larvae hindered their role as entomological evidence. Cytogenetics would aid in the identification of the immatures and their sex; the differentiation between adult species; and/ or the categorization of the taxa. Sarcophagidae received little attention concerning their karyotype measurements, especially in Egypt where there is only one unpublished thesis (El-Bassiony 2001). This mini review listed the previous studies and findings on sarcophagid’s karyotypes, especially the solitary data from Egypt.
Keywords
Entomotoxicology; Forensic entomology; Wohlfahrtia; Sarcophaga
Introduction
The study of succession pattern of insects on carrion was the most important application of forensic entomology. Family Sarcophagidae enclosed over 2500 species [1] which fed on carrion including human bodies. Additionally, sarcophagid flies have considerable forensic importance to estimate PMImini [2,3]. While flies belonging to family Calliphoridae were known as the first insect’s colonizers on carrion in most previous studies [4], other studies described Sarcophagidae as the initial and dominant colonizers on carrion in Egypt [5]. Calliphoridae was considered a key group for forensic entomologists due to the extensive biological, taxonomical, cytotaxonomical, and behavioral data, in contrary to Sarcophagidae which faced the lack of corresponding studies. The sarcophagid’s identification was difficult and perplexing, as they were cryptic and/or morphologically similar.
Main Body
The diploid number of chromosomes in most sarcophagid species, was previously noted as 2n=12 [6-15]. However, 2n=19-20 and 2n=14 have been recorded for Pseudosarcophaga affinis [9], and Helicophagella melanura [16], respectively. The karyotypes of four sarcophagid flies, Sarcophaga surcoufi (Villeneuve), Sarcophaga dux (Thomson), Sarcophaga aegyptiaca (Salem), and Wohlfahrtia nuba (Wiedemann), were studied from three locations in Egypt [14], and W. nuba has reduced 2n=10 number of chromosomes (Table 1, Figure 1). This case has never been reported before in Sarcophagidae, except for Peckia (Squamatodes) ingens [17], but this case was found in many calyptrate Diptera such as Muscina stabulans (Muscidae), Haematobia irritans (Muscidae) [18], Stomoxys calcitrans (Muscidae) [19], and Cephalopina titillator (Oestridae) [20]. Differences in both chromosome size and number can result from fissions and fusions [21]. Chromosomal fusion, especially those which decreased the chromosome number, was explained lately as a trend of karyotype evolution in many organisms [22]. Also, evolutionary turnover can convert autosomes to sex chromosomes and revert them back through the species diverge [23,24].
|
Species |
Chromosome number |
TCL (µm) |
||||||
|
I |
II |
III |
IV |
V |
X |
Y |
||
|
Parasarcophaga argyrostoma [11] Arm ratio Designation |
1.15 m |
1.81 sm |
1.82 sm |
1.17 m |
1.24 m |
NA t |
NA (15)* t |
NA |
|
Parasarcophaga argyrostoma [12] length Arm ratio Designation |
1.19 m |
1.82 sm |
1.89 sm |
1.18 m |
1.33 m |
4.9±0.24 NA NA |
3.2(7~10)* NA NA |
48.8 ±2.80 |
|
Parasarcophaga misera [12] length Arm ratio Designation |
1.67 m |
1.97 sm |
2.01 sm |
1.31 m |
1.44 m |
0.9±0.06 NA NA |
0.5(5~10)* NA NA |
48.9 ±3.11 |
|
Parasarcophaga albiceps [12] length Arm ratio Designation |
1.17 m |
1.19 m |
1.09 m |
1.16 m |
1.63 m |
1.2±0.01 NA NA |
NA NA NA |
51.5 ±1.27 |
|
Parasarcophaga ruficornis [12] length Arm ratio Designation |
1.44 m |
1.39 m |
1.57 m |
1.24 m |
1.19 m |
8.9±0.55 NA NA |
6.3(5~10)* NA NA |
64.3 ±1.19 |
|
Parasarcophaga knabi [12] length Arm ratio Designation |
1.18 m |
1.20 m |
1.17 m |
1.15 m |
1.19 m |
13.3±0.53 1.28 m |
9.7(10)* 1.15 m |
48.9 ±2.10 |
|
Pattonella intermutans [13] length Arm ratio Designation |
5.3 1.23 m |
4.5 1.91 sm |
3.9 1.45 m |
3.8 1.75 sm |
2.9 1.16 m |
7.7 3.32 st |
3.4(10)* 2.1 sm |
NA
|
|
Sarcophaga surcoufi [14] Length ± S.E. Arm ratio Designation |
6.06± 0.17 1.67 m |
5.68 ± 0.16 1.15 m |
5.31 ± 0.15 1.74 sm |
5.01 ± 0.17 1.83 sm |
4.66 ± 0.14 1.15 m |
5.83±0.21 1.17 m |
4.53 ± 0.19(23~50)* 1.28 m |
26.70 ± 0.74 |
|
Sarcophaga dux [14] length ± S.E. Arm ratio Designation |
5.00 ± 0.19 1.45 m |
4.77 ± 0.16 1.22 m |
4.49 ± 0.15 1.74 sm |
4.27±0.12 1.75 sm |
4.01±0.12 1.23 m |
5.07 ± 0.20 1.33 m |
3.77 ± 0.14(25~50)* 1.50 m |
22.52 ± 0.72 |
|
Sarcophaga aegyptiaca [14] Length ± S.E. Arm ratio Designation |
5.39 ± 0.20 1.52 m |
4.91 ± 0.20 1.26 m |
4.63 ± 0.18 1.79 sm |
4.30 ± 0.17 1.65 m |
4.08 ± 0.17 1.28 m |
4.82 ± 0.20 1.47 m |
3.80 ± 0.14(21~50)* 1.39 m |
23.24 ± 0.90 |
|
Wohlfahrtia nuba [14] Length ± S.E. Arm ratio Designation |
4.17 ± 0.12 1.57 m |
3.76 ± 0.09 1.59 m |
3.50 ± 0.06 1.32 m |
3.22 ± 0.10 1.39 m |
------- ------- ------- |
4.81±0.07 2.06 sm |
3.58 ± 0.07(23~50)* ------ t |
14.63 ± 0.25 |
Table 1: Mean length (µm), arm ratio and designation of chromosomes of somepublished and unpublished data of sarcophagid flies’ karyotypes [1-14].
TCL=mean total haploid autosomal complement length,
NA=not applicable,
m=metacentric chromosome,
sm=submetacentric chromosome,
st=subtelocentric, and t= telocentric chromosome.
*Number of Y chromosomes measured~ to the total number of chromosomes measured, in each species, in brackets.
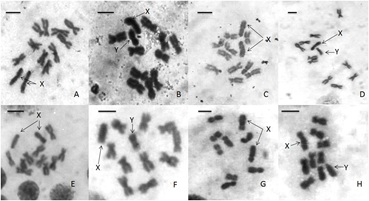 Figure 1: Neuroganglial chromosomes of four sarcophagid species [14].
Figure 1: Neuroganglial chromosomes of four sarcophagid species [14].
A: Female and B: male Sarcophaga surcoufi,
C: Female
D: male Sarcophaga dux,
E: Female
F: male Sarcophaga aegyptiaca
G: Female and H: male Wohlfahrtia nuba. Arrows show X and Y chromosomes. Bar=5μm.
Several authors described the differences between the designations of autosomal pairs and sex pair in several Sarcophagidae (Table 1) [9,10,13,14]. The sex pair showed XX designation in females, and XY in males [25]. Three metacentric and two submetacentric autosomal pairs were observed in Pattonella intermutans (Sarcophagidae), and also the X and Y chromosomes were described as subtelocentric and submetacentric respectively [13]. In the four Egyptian species (Figure 1), both S. surcoufi and S. dux have four metacentric pairs of chromosomes and two submetacentric chromosomes [14]. On the other hand, S. aegyptiaca chromosomes were all metacentric except for the third autosome. In the same study, the autosomes of W. nuba have metacentric designation, while X and Y chromosomes were submetacentric and telocentric, respectively [14]. While the autosomal morphology of Sarcophagidae was more or less similar in the different species (Table 1), the sex chromosomes were heterochromosomes and species specific [10]. The sex chromosomes were greatly varied in size and morphology among species within several genera. Nine sarcophagid flies have acrocentric sex chromosomes [9], three Parasarcophaga species have metacentric and telocentric ones [12], two Parasarcophaga [12], and nine Boettcherisca species [15] have small dot-like chromosomes. The large verity in the size and morphology of the sex chromosomes might be a result of heterochromatin accumulation or deletion, where the changes in the amount and distribution of heterochromatin resulted mainly in the evolution of the karyotype of a species [12,26].
The previous studies on genome size evaluated the phylogenetic relations in the taxonomy field [27-29]; estimated the completeness of genomes [30]; and helped in assessing genetic linkage maps [30]. Genome size was usually considered constant within species, and could vary widely between closely related species [22]. Limited intraspecific variation was an assumption in measurement and comparison of genome sizes. The inter- and intraspecific genome size variation might be due to differences in heterochromatin content and the amount of repetitive DNA in the autosomal complement [31]. The average Total haploid Complement Lengths (TCL) of species belonging to the genera Pseudosarcophaga, Sarcophaga, Kellymyia and Protodexia were recorded in 1953 [9]. Also, the karyotype measurements of S. surcoufi exhibit significantly longer chromosome lengths, and longer TCL than S. dux, S. aegyptiaca and Wohlfahrtia nuba [14].
Conclusion
The genetics analyses could provide cue for the chromosome’s evolution in sarcophagid flies, and could supply information which is of both forensic significance and evolutionary interest.
Funding
Not applicable
Acknowledgements
The author would like to express her gratitude to Dr. Mahitab Ezzat El Daly for reviewing the English language.
References
- Pape T (1996) Catalogue of the Sarcophagidae of the world (Insecta; Diptera). Mem Entomol Int 8: 1-558.
- Amendt J, Zehner R, Krettek R (2004) Forensic entomology. Naturwissenschaften 91: 51-65.
- Byrd JH, Castner JL (2009) Forensic entomology: The utility of arthropods in legal investigations. Boca Raton: CRC Press.
- Campobasso CP, Di Vella G, Introna F (2001) Factors affecting decomposition and Diptera colonization. Forensic Sci Int 120: 18-27.
- El-Bassiony GM (2020) Blowflies (Diptera: Calliphoridae) As Forensic Indicators in Egypt with Special Reference to the Development Data of Lucilia Cuprina (Wiedemann). Sumerianz J Biotechnol 3: 43-51.
- Stevens NM (1908) A study of the germ cells of certain Diptera, with reference to the heterochromosomes and the phenomena of synapsis. Jour Exptl Zool 5: 359-374.
- Metz CW (1916) Chromosome studies of the Diptera. II. The paired association of chromosomes in the Diptera, and its significance. Jour Exptl Zool 21: 213-280.
- Keuneke W (1924) Über die spermatogenese einiger Dipteren. Z Zell Gewebelehre 1: 357-412.
- Boyes JW (1953) The chromosomes of the higher Diptera. II. Differentiation of the sarcophagid species. Canadian Jour Zool 31: 561-576.
- Boyes JW, Brink JMV (1965) Chromosomes of calyptrate Diptera.Canad J Genet Cytol 7: 537-550.
- Kaul D, Tewari RR (1977) polytene chromosomes of the flesh-fly Parasarcophaga argyrostoma (Sarcophagidae: Diptera). Genetica 49: 189-197.
- Kaul D, Chaturvedi R, Gaur P, Tewari RR (1978) Cytogenetics of the genus Parasarcophaga (Sarcophagidae: Diptera). Chromosoma 68: 73-82.
- Parise-Maltempi PP, Avancini RMP (2000) Cytogenetics of the neotropical flesh fly Pattonella intermutans (Diptera, Sarcophagidae). Genetics and Molecular Biology 23: 563-567.
- El-Bassiony GM (2001) Cytogenetic and biochemical genetic investigation of four sarcophagid flies (Diptera: Sarcophagidae). Ph.D. thesis supervised by: FK Adham, ME Zowail, AH Omar.
- Agrawal UR, Bajpai N, Tewari RR, Kurahadhi H (2010) Cytogenetics of Flesh Flies of the Genus Boettcherisca (Sarcophagidae: Diptera). Cytologia 75: 149-155.
- Tewari RR, Kurahashi H (1985) The karyotype of Helicophagella melanura (Meigen) (Sarcophagidae: Diptera). CIS 38: 14-15.
- Nunes GM (2018) Citogenética, citotaxonomia e cariossistemática de moscas de importância forense. Master thesis.
- Boyes JW (1958) Chromosomes in classification of Diptera. Proc Tenth Internatl Cong Entomol 2: 899-906.
- LaChance LE (1964) Chromosome studies in three species of Diptera (Muscidae and Hypodermatidae). Ann Entomol Soc Am 57: 69-73.
- El Bassiony GM (2004) Karyotype and nucleic acids patterns of the camel nasal bot fly Cephalopina titillator. Bull Ent Soc Egypt 81: 129-139.
- White MJD (1973) Animal Cytology and Evolution. Cambridge University Press. Cambridge 961.
- Gokhman VE, Kuhn KL, Woolley JB, Hopper KR (2017) Variation in genome size and karyotype among closely related aphid parasitoids (Hymenoptera, Aphelinidae). Comp Cytogenet 11: 97-117.
- Larracuente, AM, Noor MA, Clark AG (2010) Translocation of Y-linked genes to the dot chromosome in Drosophila pseudoobscura. Mol Biol Evol 27: 1612-1620.
- Bachtrog D, Mank JE, Peichel CL, Kirkpatrick M, Otto SP, et al. (2014) Sex Determination: Why So Many Ways of Doing It? PLoS Biol 12: e1001899.
- Rashmi S, Pratima G (2015) Revelation of heterochromatin heterogeneity in Sarcophagid chromosomes using DNA ligand Mithramycin. Caryologia 68: 55-60.
- Hanrahan SJ, Johnston JS (2011) New genome size estimates of 134 species of arthropods. Chromosome Res 19: 809-823.
- Alekseeva SS, Andreeva YV, Wasserlauf IE, Sibataev AK, Stegniy VN (2020) Analysis of the Metaphase Chromosome Karyotypes in Imaginal Discs of Aedes communis, punctor, Ae. intrudens, and Ae. rossicus (Diptera: Culicidae) Mosquitoes. Insects 11: 63.
- Ivorra T, Martínez-Sánchez A, Rojo S (2021) Review of Synthesiomyia nudiseta(Diptera: Muscidae) as a useful tool in forensic entomology. Int J Legal Med 135: 2003-2015.
- Gregory TR, Johnston JS (2008) Genome size diversity in the family Drosophilidae. Heredity 101: 228-238.
- Gokhman VE, Kuznetsova VG (2006) Comparative insect karyology: Current state and applications. Entomol Rev 86: 352-368.
- Biemont C (2008) Within-species variation in genome size. Heredity 101: 297-298.
Citation: El-Bassiony GM (2022) The Karyotype of the Forensically Important Flesh Flies (Diptera: Sarcophagidae), Mini Review. Forensic Leg Investig Sci 8: 069.
Copyright: © 2022 Ghada Mohamed El-Bassiony, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.

